Introduction
CRISPR-based genome editing has transformed modern biology, enabling precise and programmable manipulation of DNA across virtually all organisms. Originally derived from a bacterial immune system, CRISPR technologies have rapidly evolved from simple gene disruption tools into sophisticated platforms capable of rewriting genetic information with unprecedented precision. ([Synthego][1])
Over just a decade, CRISPR has moved from discovery to clinical application, culminating in the first approved CRISPR-based therapy for genetic disease—marking a turning point in genomic medicine. ([Innovative Genomics Institute (IGI)][2])
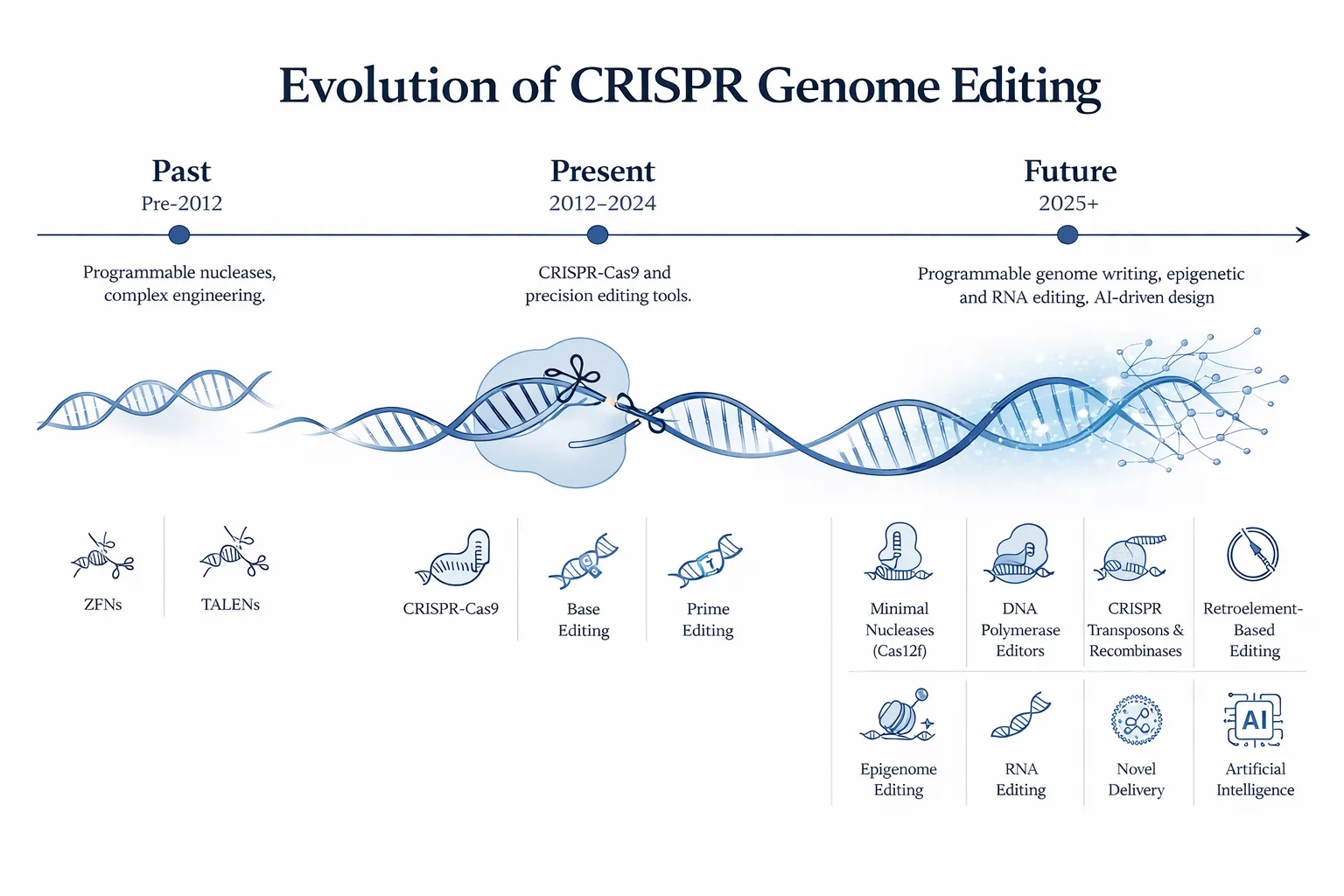
1. The Evolution of CRISPR Technologies
1.1 Early Genome Editing Foundations (Pre-CRISPR)
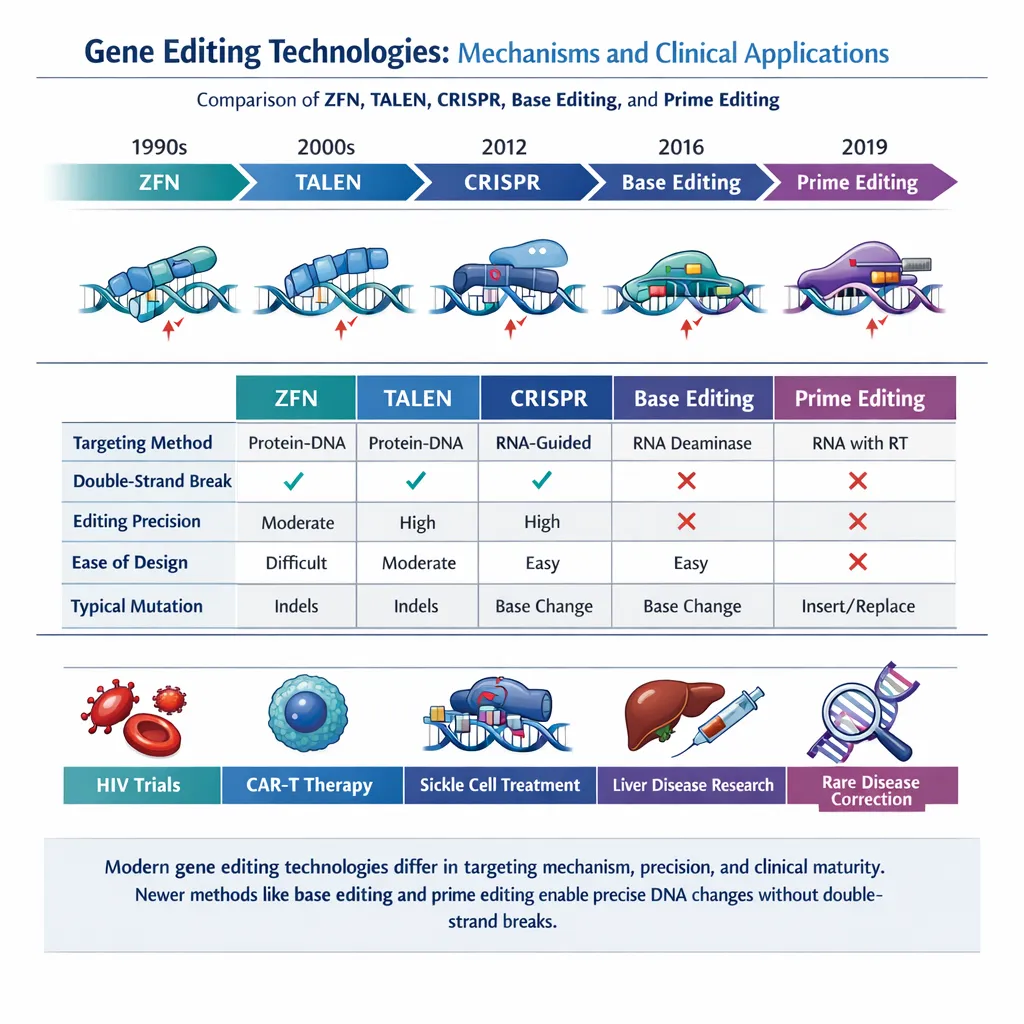
Before CRISPR, genome editing relied on tools such as:
- Zinc Finger Nucleases (ZFNs)
- TALENs (Transcription Activator-Like Effector Nucleases)
These systems required complex protein engineering for each target site, limiting scalability and adoption.
CRISPR revolutionized the field by introducing:
- RNA-guided DNA targeting
- Simple and flexible design
- High editing efficiency
Originally discovered as a bacterial immune system, CRISPR-Cas systems protect against viral infection by recognizing and cutting foreign DNA.
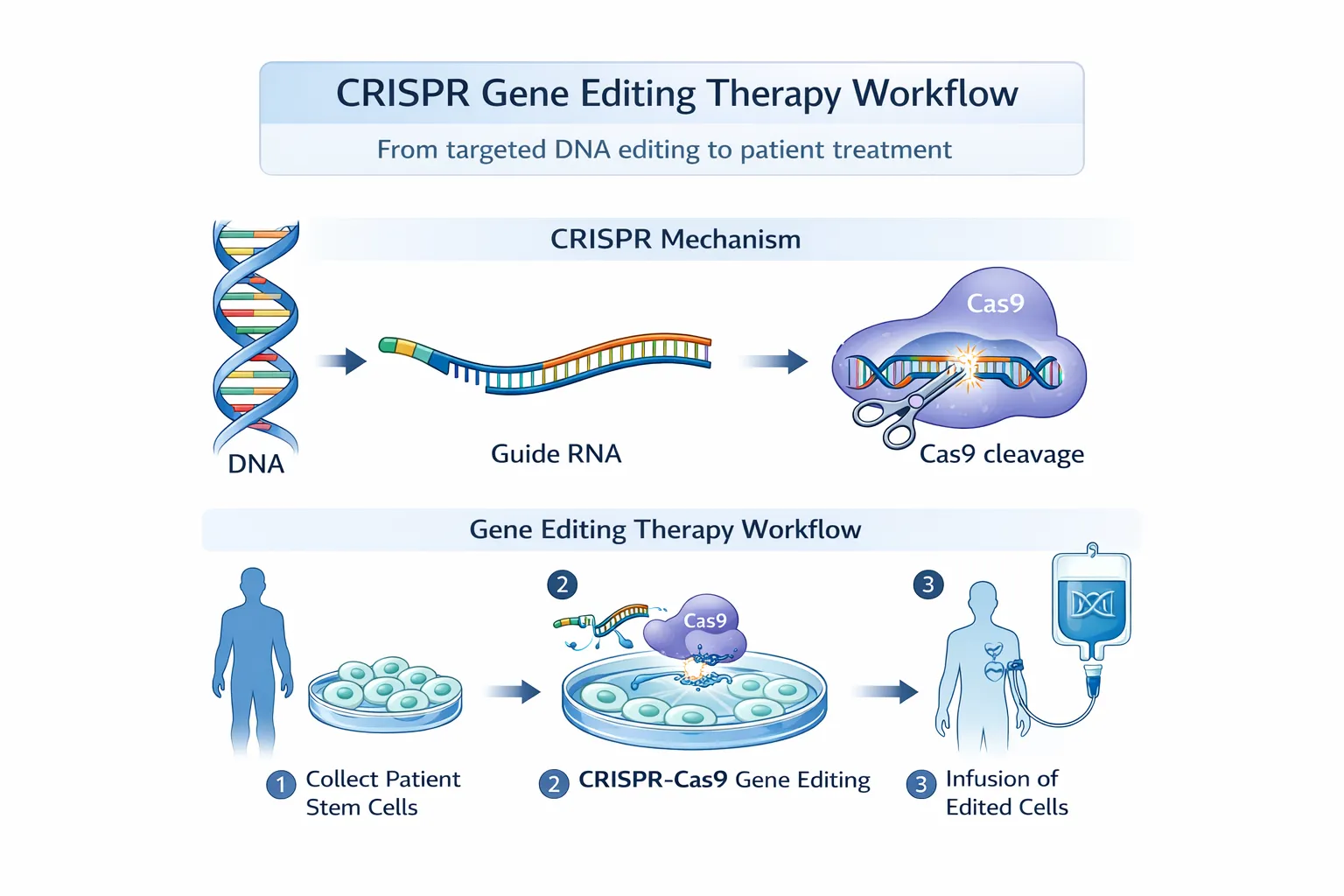
1.2 The CRISPR-Cas9 Revolution (2012–Present)
CRISPR-Cas9 introduced a paradigm shift by replacing protein engineering with RNA-guided targeting. A guide RNA (gRNA) directs the Cas9 nuclease to a specific DNA sequence, where it introduces a double-strand break. ([Synthego][1])
This break is repaired by:
- Non-homologous end joining (NHEJ): introduces insertions/deletions (knockouts)
- Homology-directed repair (HDR): enables precise edits
CRISPR’s simplicity, efficiency, and programmability rapidly made it the dominant genome editing platform. ([Stanford News][3])
2. Current CRISPR Technologies (Deep Dive)
Modern CRISPR systems extend far beyond simple DNA cutting.
2.1 CRISPR Nucleases (Cas9, Cas12, Cas13)
- Cas9: Most widely used; introduces double-strand DNA breaks
- Cas12a (Cpf1): Offers staggered cuts and multiplex editing potential
- Cas13: Targets RNA instead of DNA
These systems enable gene knockout, activation, repression, and RNA targeting.
2.2 Base Editing
Base editors enable single-nucleotide changes without double-strand breaks, significantly reducing unintended mutations.
Types:
- Cytosine Base Editors (C→T)
- Adenine Base Editors (A→G)
These tools are especially valuable for correcting point mutations, which account for a large fraction of genetic diseases. ([Labiotech.eu][4])
2.3 Prime Editing
Prime editing is a “search-and-replace” genome editing system capable of:
- Insertions
- Deletions
- Substitutions
It uses a Cas9 nickase fused to reverse transcriptase and a prime editing guide RNA (pegRNA), enabling highly versatile and precise editing without double-strand breaks. ([Labiotech.eu][4])
2.4 Epigenome Editing
CRISPR systems can be repurposed to modify gene expression without altering DNA sequence, by:
- Recruiting transcriptional activators/repressors
- Modifying chromatin states
This allows reversible control of gene function and is critical for studying regulatory biology. ([Stanford News][3])
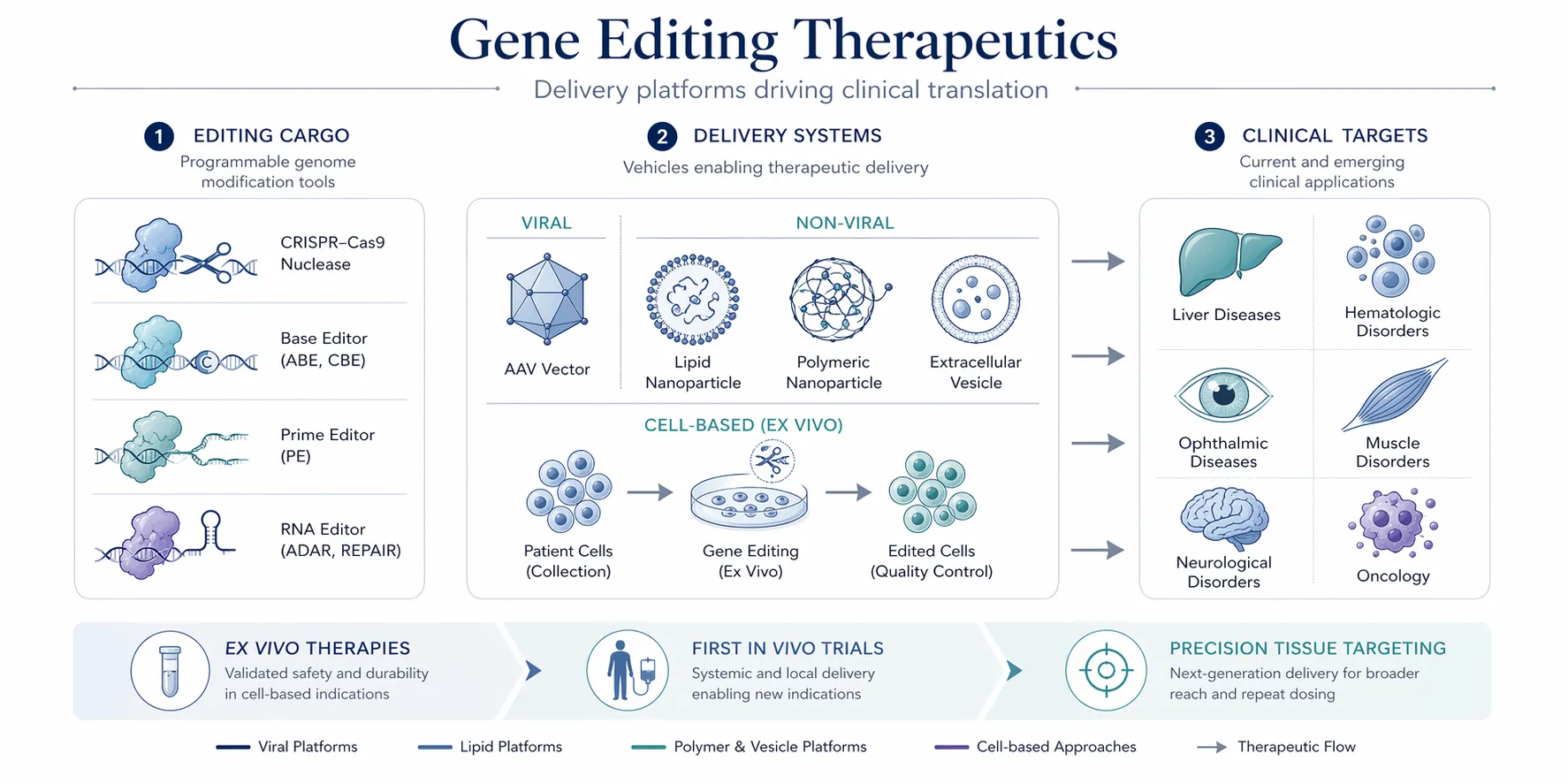
2.5 Delivery Modalities
A key challenge in CRISPR applications is delivery:
- Ex vivo editing: Cells edited outside the body and reintroduced
- In vivo editing: Direct delivery into tissues
Delivery systems include viral vectors, lipid nanoparticles, and emerging non-viral platforms. ([canjhealthtechnol.ca][5])
Comparative Table of CRISPR Technologies
| Technology | Mechanism | Molecular Target | Edit Type | Key Advantages | Limitations |
|---|---|---|---|---|---|
| CRISPR-Cas9 | Double-strand break | DNA | Knockout / HDR edits | Simple, versatile | Off-target effects, DSB toxicity |
| Cas12a | Staggered DNA cuts | DNA | Multiplex editing | Improved specificity | Less mature ecosystem |
| Cas13 | RNA targeting | RNA | Transcript knockdown/editing | No permanent DNA changes | Transient effects |
| Base Editing | Deamination | DNA | Single-base changes | No DSB, high precision | Limited to specific mutations |
| Prime Editing | Reverse transcription | DNA | Insertions/deletions/substitutions | Highly versatile | Lower efficiency, complexity |
| Epigenome Editing | Transcription modulation | DNA/chromatin | Gene regulation | Reversible, non-destructive | Temporary effects |
| Future Multiplex Editors | Multi-site targeting | DNA/RNA | Complex edits | Systems-level editing | Delivery challenges |
🚀 3. Future CRISPR Technologies
Emerging CRISPR technologies are addressing current limitations such as inefficient large DNA insertion, delivery constraints, and off-target effects. These next-generation systems move beyond simple editing toward programmable genome writing, regulation, and system-level engineering.
3.1 Minimal Ancestral RNA-Guided Nucleases
One major limitation of current CRISPR systems (e.g., Cas9) is their relatively large size, which complicates delivery—especially via viral vectors like AAV.
Recent efforts have focused on discovering and engineering minimal or ancestral RNA-guided nucleases, such as Cas12f (≈400–700 amino acids):
- Derived from compact CRISPR systems found in bacteria
- Significantly smaller than Cas9 (~1,300 aa)
- Enable easier packaging into delivery systems
These nucleases:
- Retain RNA-guided targeting
- Can be engineered for higher activity via structure-guided design and mutational scanning
- Serve as scaffolds for fusion systems (e.g., base editors, transcriptional regulators)
👉 These systems are expected to become critical for in vivo gene therapy applications due to improved delivery efficiency. ([ScienceDirect][1])
3.2 DNA Polymerase Editors
DNA polymerase-based editing systems represent a new class of template-driven genome editing technologies.
Example:
- Fusion of Cas9 nickase (nCas9) with a DNA polymerase
Mechanism:
- Target DNA is nicked (single-strand cut)
- A DNA polymerase extends the strand using a synthetic template
- The new sequence is incorporated during repair
Key features:
- Enables continuous DNA synthesis at target sites
- Expands editing window beyond base editing
- Reduces reliance on double-strand breaks
👉 These “writing-based editors” move CRISPR from cutting tools to DNA synthesis platforms. ([PMC][2])
3.3 CRISPR-Guided Recombinases and Transposons
One of the biggest unmet needs in genome editing is efficient insertion of large DNA fragments (e.g., full genes).
CRISPR-Associated Transposons (CAST)
- Combine CRISPR targeting with transposase enzymes
- Enable RNA-guided insertion of large DNA sequences
- Do not rely on host DNA repair mechanisms
Mechanism:
- CRISPR system identifies genomic target
- Transposase inserts DNA payload at that site
CRISPR-Guided Recombinases
- Use site-specific recombinases (e.g., Bxb1)
- CRISPR introduces recombination sites (attB/attP)
- Recombinase integrates large DNA sequences
Emerging platforms:
- PASTE (Programmable Addition via Site-specific Targeting Elements)
- PASSIGE systems
👉 These technologies enable gene-scale insertions, overcoming major HDR limitations. ([ScienceDirect][1])
3.4 Retroelement-Based Editing
Retroelement-based editing leverages mechanisms from retrotransposons and reverse transcription.
Mechanism:
- Cas9 nickase generates a single-strand break
- A reverse transcriptase synthesizes DNA from an RNA template
- The new DNA is inserted at the target site
This process—known as target-primed reverse transcription (TPRT)—enables:
- Integration of large DNA sequences
- Editing independent of double-strand breaks
- Potential for editing in non-dividing cells
👉 This approach is particularly promising for neuronal and post-mitotic tissues, where HDR is inefficient. ([ScienceDirect][1])
3.5 Epigenome Editing (Next-Generation)
Future epigenome editing expands beyond current CRISPRi/a systems toward stable, heritable, and programmable chromatin control.
Mechanism:
Use dCas9 (catalytically inactive Cas9) fused to:
DNA methyltransferases → gene silencing (CRISPRoff)
Demethylases / activators → gene activation (CRISPRon)
Key advances:
- Heritable gene silencing without DNA sequence change
- Reversible gene activation systems
- Multi-locus epigenetic reprogramming
👉 Enables cell fate engineering and disease reprogramming without permanent genome edits. ([ScienceDirect][1])
3.6 RNA Editing Technologies
RNA-targeting CRISPR systems (e.g., Cas13) are evolving into precision RNA editing platforms.
Capabilities:
- Transient modification of gene expression
- Correction of RNA transcripts without DNA alteration
- Programmable RNA degradation or editing
Advantages:
- No permanent genomic change
- Reduced safety concerns
- Ideal for reversible therapies
Emerging tools:
- RNA base editors (A→I editing)
- RNA-guided transcript modulation
👉 RNA editing represents a safer and reversible alternative for therapeutic interventions. ([ScienceDirect][1])
3.7 Novel Delivery Methods
Delivery remains a major bottleneck in CRISPR translation.
Emerging Delivery Technologies:
1. Lipid Nanoparticles (LNPs)
- Non-viral delivery
- Widely used in mRNA therapeutics
2. Engineered Viral Vectors
- AAV variants with improved tissue specificity
- Reduced immunogenicity
3. Virus-Like Particles (VLPs)
- Deliver CRISPR ribonucleoproteins (RNPs)
- Transient expression reduces off-target effects
4. Physical Delivery Methods
- Electroporation
- Microinjection
Future directions:
- Tissue-specific targeting
- Programmable delivery systems
- Multi-component delivery (editor + template)
👉 Advances in delivery will determine the clinical scalability of CRISPR therapies.
3.8 Artificial Intelligence in CRISPR
AI is becoming a central driver of next-generation genome editing.
Key Applications:
1. Guide RNA Design
- Predicts optimal gRNA sequences
- Minimizes off-target effects
2. Protein Engineering
- Designs improved Cas variants
- Enhances specificity and efficiency
3. Editing Outcome Prediction
- Models DNA repair outcomes
- Improves editing precision
4. Multi-omics Integration
- Links genome editing with transcriptomic and epigenomic data
👉 AI enables data-driven, predictive genome engineering, accelerating discovery and therapeutic development. ([ScienceDirect][1])
🔮 Summary of Future Directions
| Category | Innovation | Impact |
|---|---|---|
| Compact nucleases | Cas12f, minimal systems | Improved delivery |
| DNA writing | Polymerase editors | Continuous DNA synthesis |
| Large insertions | Transposons, recombinases | Gene-scale editing |
| Reverse transcription | Retroelement editing | Editing in non-dividing cells |
| Regulation | Epigenome editing | Reversible gene control |
| RNA editing | Cas13-based tools | Transient therapies |
| Delivery | LNPs, VLPs | Clinical translation |
| AI | Predictive modeling | Precision and scalability |
5. Current Applications in Basic Research and Human Medicine
5.1 Basic Research
CRISPR has revolutionized research by enabling:
- Functional genomics screens
- Disease modeling
- Gene regulatory network mapping
It allows rapid and scalable interrogation of gene function across biological systems.
5.2 Human Medicine
Approved Therapies
The first CRISPR-based therapy (Casgevy) has been approved for:
- Sickle cell disease
- Beta-thalassemia ([Innovative Genomics Institute (IGI)][2])
Therapeutic Areas
- Genetic diseases: Correction of pathogenic mutations
- Cancer: Engineering CAR-T cells
- Infectious diseases: Targeting viral genomes
- Autoimmune disorders: Immune system modulation
CRISPR enables treatment at the root genetic cause, rather than managing symptoms. ([PMC][7])
6. Challenges and Limitations
Despite rapid progress, several challenges remain:
- Off-target editing
- Delivery efficiency
- Immune responses
- Ethical considerations
- High cost and scalability
Ongoing innovations are actively addressing these limitations. ([PubMed][8])
7. Future Outlook
CRISPR technologies are evolving toward:
- Precision medicine at scale
- Multi-gene and pathway-level editing
- Integration with AI and multi-omics
- Real-time and reversible editing systems
These advances will likely redefine how we treat disease and engineer biology in the coming decades.
References
- What is CRISPR: Your Ultimate Guide 🔗
- CRISPR Clinical Trials: A 2024 Update - Innovative Genomics Institute 🔗
- What is CRISPR? A bioengineer explains - Stanford Report 🔗
- CRISPR-Cas9: The gene editing tool changing the world 🔗
- CRISPR Technologies for In Vivo and Ex Vivo Gene Editing 🔗
- New CRISPR tool enables more seamless gene editing 🔗
- Recent applications, future perspectives, and limitations of the CRISPR-Cas system 🔗
- Past, present, and future of CRISPR genome editing technologies 🔗