Introduction
The convergence of deep learning and multi-omics data is transforming how we understand biology and disease. Modern biomedical research now generates vast amounts of heterogeneous data—from genomics and transcriptomics to proteomics, metabolomics, and imaging. Individually, each modality offers partial insight. Together, they provide a holistic view of biological systems.
Deep learning has emerged as a powerful approach to integrate these diverse datasets, enabling breakthroughs in disease prediction, patient stratification, and biomarker discovery.
Deep learning enables multi-omics integration by combining genomics, transcriptomics, proteomics, and clinical data into unified models. But how do Deep Learning approaches in AI compare to each other?
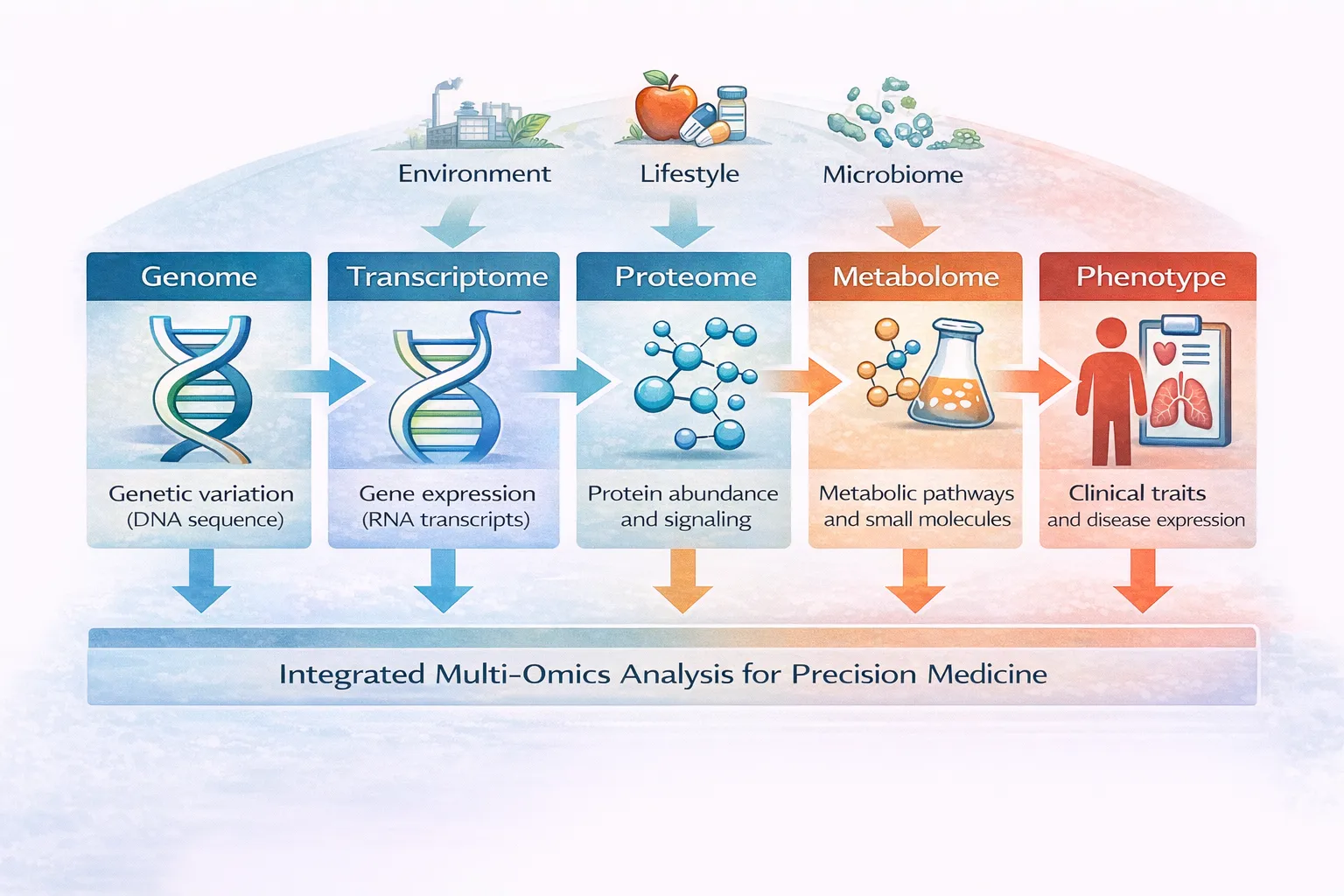
What Is Multi-Omics Data Integration?
Multi-omics refers to the simultaneous analysis of multiple biological data layers, including:
- Genomics (DNA)
- Transcriptomics (RNA expression)
- Proteomics (protein levels)
- Epigenomics (gene regulation)
- Metabolomics (metabolic processes)
These datasets are complementary, capturing different aspects of biological function. Integrating them allows researchers to uncover complex interactions that are not visible from a single modality alone.
Why Deep Learning?
Traditional statistical and machine learning approaches struggle with multi-omics data due to:
- High dimensionality (thousands of features)
- Heterogeneity across data types
- Missing or incomplete datasets
Deep learning addresses these challenges by learning nonlinear relationships and transforming diverse inputs into shared latent representations, enabling effective integration.
Core Deep Learning Approaches for Multi-Omics Integration
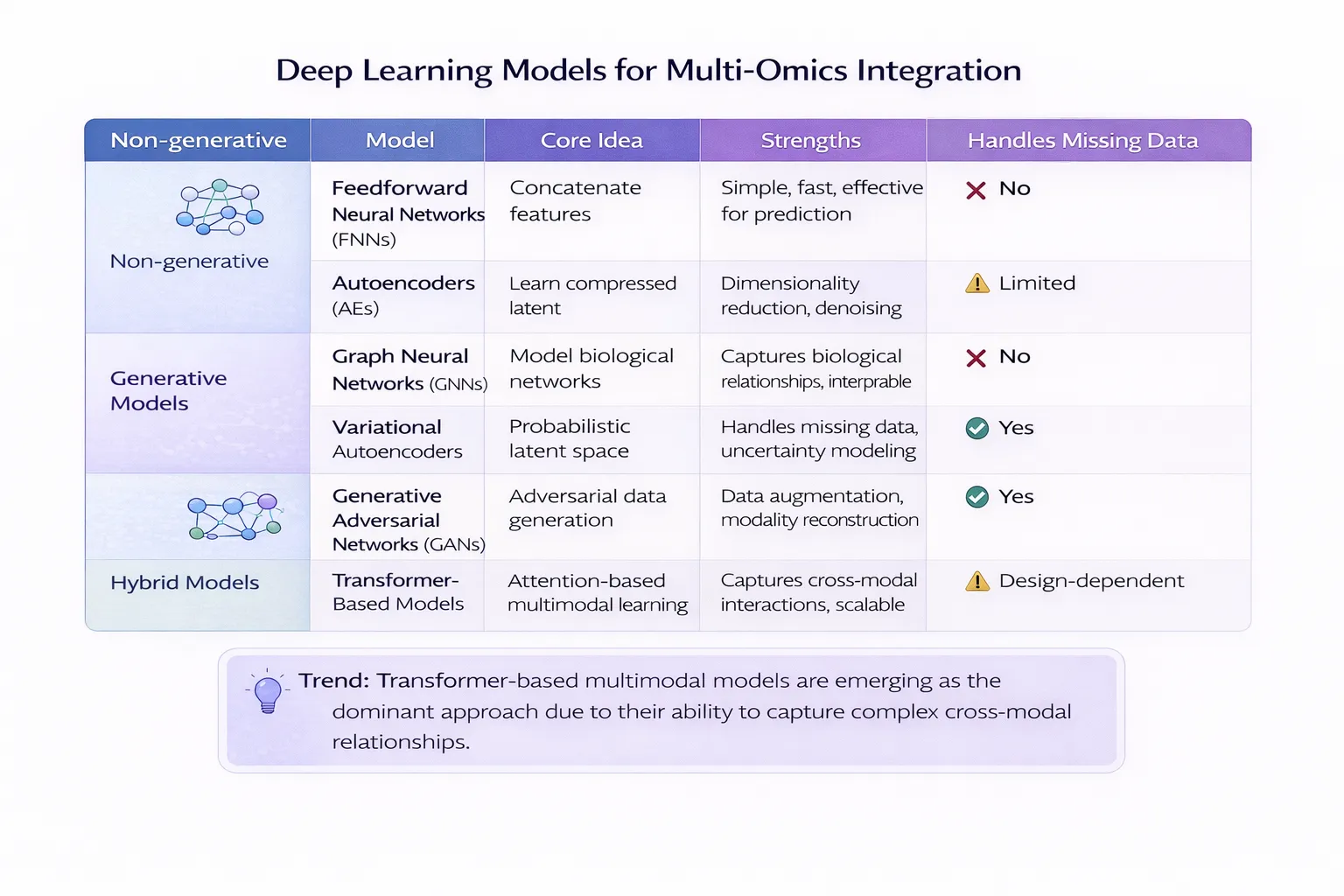
Deep learning methods for multi-omics integration can be broadly categorized into non-generative (discriminative) and generative approaches. Each class differs in how it learns representations, handles missing data, and supports downstream biological interpretation.
1. Feedforward Neural Networks (FNNs)
Feedforward Neural Networks are among the simplest deep learning approaches for multi-omics integration. In this setup, features from different omics layers are typically concatenated into a single input vector, which is then passed through multiple dense layers to learn predictive patterns. This approach is often used for classification tasks such as disease prediction or subtype identification.
Strengths
- Simple and easy to implement
- Works well with well-curated, complete datasets
- Efficient training and low computational overhead
Weaknesses
- Ignores modality-specific structure
- Poor handling of missing data
- Limited interpretability in high-dimensional settings
2. Autoencoders (AEs)
Autoencoders are widely used for representation learning and dimensionality reduction in multi-omics data. They consist of an encoder that compresses input data into a latent space and a decoder that reconstructs the original input.
In multi-omics settings, separate encoders may be used per modality, followed by a shared latent representation.
Strengths
- Effective for high-dimensional data compression
- Can denoise noisy biological datasets
- Enables intermediate (representation-level) integration
Weaknesses
- Latent representations may lack biological interpretability
- Sensitive to hyperparameter tuning
- Struggles with missing modalities unless modified
3. Graph Neural Networks (GNNs)
Graph Neural Networks incorporate biological prior knowledge, such as gene–gene interaction networks or protein–protein interaction graphs. Nodes represent biological entities, and edges capture relationships, allowing the model to learn structured dependencies across omics layers.
Strengths
- Captures biological relationships explicitly
- Improves interpretability via network structure
- Suitable for pathway-level analysis
Weaknesses
- Requires high-quality prior biological networks
- Computationally intensive for large graphs
- Difficult to scale with many modalities
4. Variational Autoencoders (VAEs)
Variational Autoencoders extend traditional autoencoders by learning a probabilistic latent space, enabling the model to capture uncertainty and variability in biological data. VAEs are particularly useful for data integration, imputation, and simulation in multi-omics studies.
Strengths
- Handles missing data through probabilistic modeling
- Enables data generation and augmentation
- Captures uncertainty in biological systems
Weaknesses
- More complex training than standard autoencoders
- Risk of oversmoothing latent representations
- Requires careful tuning of loss functions
5. Generative Adversarial Networks (GANs)
GANs consist of two competing networks—a generator and a discriminator—that learn to produce realistic synthetic data. In multi-omics integration, GANs can be used to generate missing modalities, augment datasets, or align distributions across omics layers.
Strengths
- Powerful for data augmentation
- Can reconstruct missing modalities
- Learns complex, high-dimensional distributions
Weaknesses
- Training instability (mode collapse)
- Difficult to interpret biologically
- Requires large datasets for effective training
6. Transformer-Based Models
Transformer architectures leverage attention mechanisms to model relationships across features and modalities. In multi-omics integration, transformers enable cross-modality attention, learning how different omics layers interact within a shared representation space.
Strengths
- Captures long-range and cross-modal dependencies
- Highly flexible and scalable
- Strong performance in multimodal learning
Weaknesses
- Data-hungry (requires large datasets)
- High computational cost
- Limited interpretability without additional tools
Comparative Overview of Deep Learning Approaches
| Approach | Type | Key Idea | Strengths | Weaknesses | Best Use Case |
|---|---|---|---|---|---|
| FNN | Non-generative | Concatenate features and learn mapping | Simple, fast, effective for prediction | Ignores modality structure, poor with missing data | Baseline prediction tasks |
| Autoencoder | Non-generative | Learn compressed latent representation | Dimensionality reduction, denoising | Limited interpretability, struggles with missing data | Feature extraction |
| GNN | Non-generative | Model biological networks | Captures biological relationships, interpretable | Requires prior knowledge, computationally heavy | Pathway analysis |
| VAE | Generative | Probabilistic latent space | Handles missing data, uncertainty modeling | Complex training, tuning required | Data integration & imputation |
| GAN | Generative | Adversarial data generation | Data augmentation, modality reconstruction | Training instability, hard to interpret | Synthetic data generation |
| Transformer | Hybrid | Attention-based multimodal learning | Captures cross-modal interactions, scalable | Data-intensive, expensive | Multimodal fusion |
Integration Strategies in Multi-Omics Learning
Deep learning models integrate multi-omics data using three main strategies:
Early Integration (Feature-Level Fusion)
- Concatenates all modalities into a single input
- Simple but may ignore modality-specific structure
Intermediate Integration (Representation-Level Fusion)
- Learns separate representations per modality
- Combines them into a shared latent space
👉 This is the most widely used and effective approach
Late Integration (Decision-Level Fusion)
- Trains separate models per modality
- Combines predictions at the end
Each strategy balances trade-offs between interpretability, flexibility, and performance.
Key Challenges in Multi-Omics Integration
1. High Dimensionality
Multi-omics datasets often contain thousands of features, leading to the curse of dimensionality and overfitting risks.
2. Missing Data
Incomplete datasets are common due to experimental and practical limitations. Deep learning—especially generative models—helps impute missing modalities.
3. Paired vs. Unpaired Data
Data from different omics layers may not come from the same samples, complicating integration.
4. Heterogeneity Across Modalities
Different data types vary in scale, distribution, and meaning, requiring sophisticated alignment strategies.
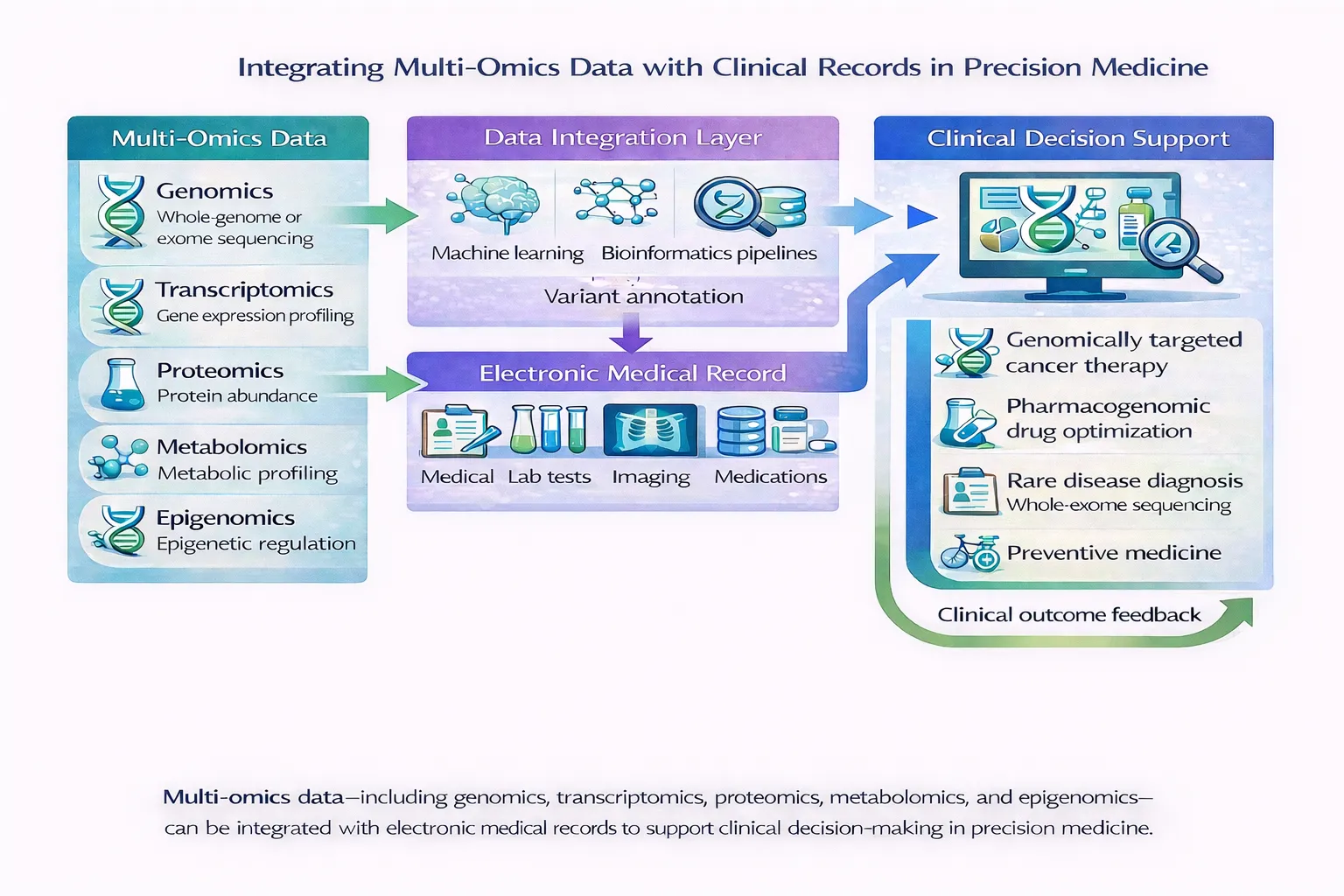
Applications in Precision Medicine
Deep learning-based multi-omics integration is enabling:
- 🔬 Disease Subtype Discovery
- 📊 Clinical Outcome Prediction
- 💊 Drug Response Modeling
- 🧬 Biomarker Discovery
These applications are central to advancing personalized and predictive healthcare.
Emerging Trends
Multimodal expansion beyond omics via integration with imaging, clinical data, and EHRs is key! See our article on cross modality integration using AI using Artificial Intelligence. In addition Large-scale pretrained models (Foundation Models & Transformers) with attention mechanisms are emerging to handle some of the challenges in this space. Improved robustness using generative and hybrid approaches could handle incomplete data which continues to be a longer term issue across healthcare space.
Conclusion
Deep learning has become a cornerstone of multi-omics data integration, enabling researchers to extract meaningful insights from complex, high-dimensional biological data.
By combining diverse data modalities into unified representations, these approaches are accelerating progress in:
- Disease understanding
- Clinical decision-making
- Precision medicine
As the field evolves—particularly with advances in **transformers and generative AI**—multi-omics integration will play an increasingly central role in the future of healthcare.
Key Takeaways
- Multi-omics data provides a comprehensive view of biological systems
- Deep learning enables integration through shared representations
- Generative models are critical for handling missing data
- Model choice depends on data, task, and interpretability needs
- Multimodal AI is driving the next wave of precision medicine