Introduction
The rapid expansion of multi-omics technologies—encompassing genomics, transcriptomics, proteomics, and metabolomics—has revolutionized biomedical research. However, these datasets are inherently high-dimensional, heterogeneous, and complex, presenting major challenges for analysis and interpretation.
An emerging and highly promising solution is the transformation of multi-omics data into image-based representations, enabling the application of advanced deep learning techniques originally developed for computer vision. This paradigm shift is unlocking new possibilities in disease classification, biomarker discovery, and precision medicine.
Why Transform Multi-Omics Data into Images?
Traditional multi-omics data is typically structured as tabular matrices. While informative, this format has limitations:
- Difficult to capture nonlinear biological relationships
- Poor scalability with high-dimensional datasets
- Limited compatibility with deep learning architectures
- Hierarchical and spatial relationships between features
- Patterns across multiple biological layers
Image-based transformation addresses these challenges by:
- Converting numerical data into spatial representations
- Enabling the use of convolutional neural networks (CNNs)
- Enhancing feature extraction and pattern recognition
This shift allows models to uncover hidden biological structures and interactions that may not be detectable using conventional methods. ([ScienceDirect][1])
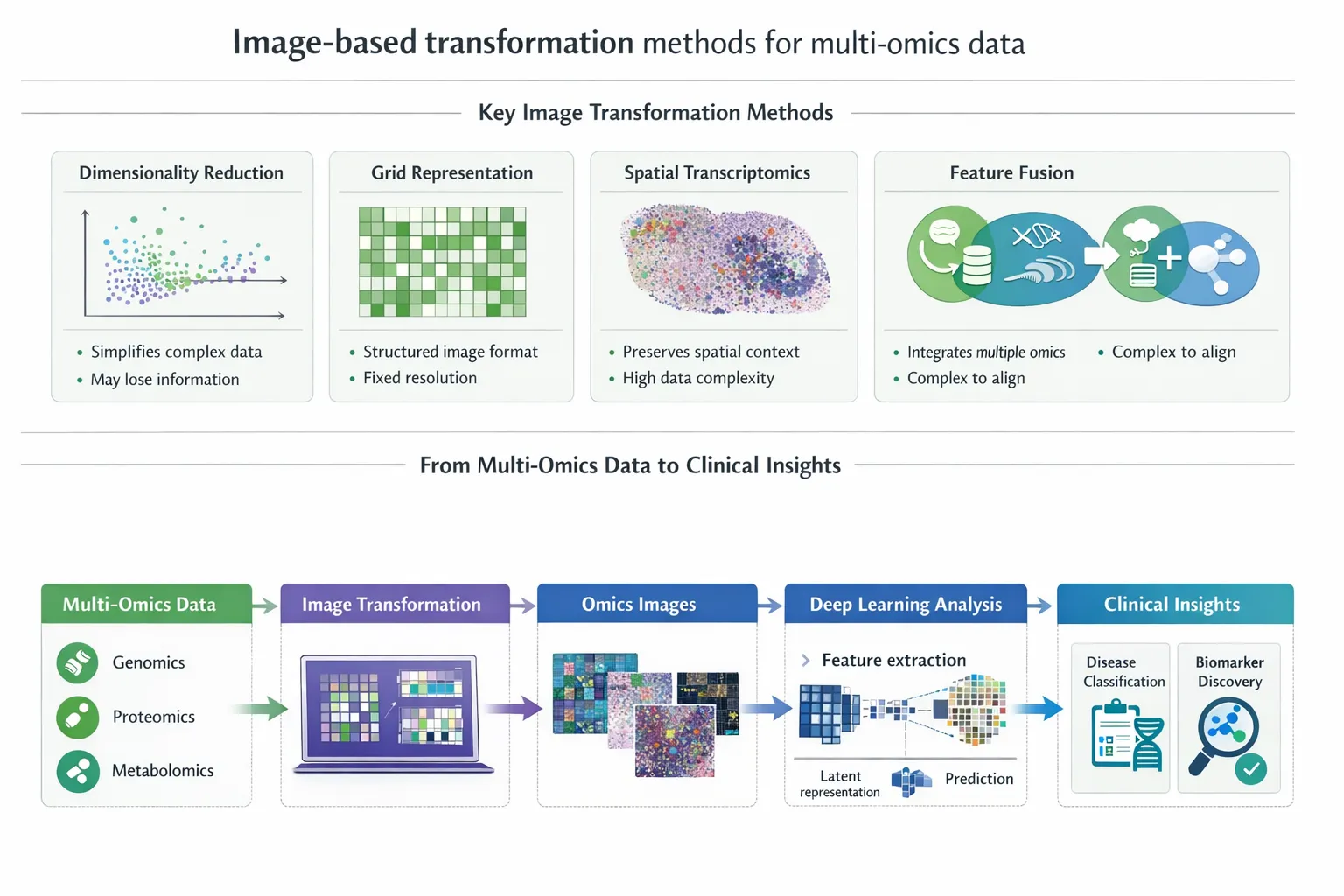
Key Image-Based Transformation Techniques
A variety of computational strategies have been developed to convert multi-omics data into image-like formats:
1. Dimensionality Reduction-Based Mapping
These methods project high-dimensional data into two-dimensional space while preserving relationships:
- t-SNE – preserves local structure
- UMAP – balances local and global structure
- Kernel PCA – captures nonlinear relationships
2. Signal Processing Transformations
These approaches treat omics data as signals:
- Fast Fourier Transform (FFT)
- Wavelet transforms
They reveal frequency-domain patterns, which may correspond to biological rhythms or regulatory signals.
3. Spatial Mapping and Layout Methods
These assign biological features to specific spatial positions:
- Chromosome-based layouts
- Network-based layouts (gene–gene interactions)
- Treemap visualizations
This preserves biological context, such as genomic proximity or pathway relationships.
4. DeepInsight and Structured Encoding Frameworks
Frameworks like DeepInsight convert tabular data into structured images optimized for deep learning.
These methods:
- Preserve feature similarity and clustering
- Enable transfer learning from image models
- Improve classification performance
Comparative Overview of Image Transformation Methods
| Method Type | Key Principle | Strengths | Limitations | Best Use Case |
|---|---|---|---|---|
| Dimensionality reduction (t-SNE, UMAP) | Projects data into 2D space | Preserves relationships, intuitive visualization | May distort global structure | Exploratory analysis, clustering |
| Signal processing (FFT, wavelets) | Converts data into frequency domain | Captures hidden patterns | Less biologically intuitive | Time-series or dynamic omics |
| Spatial mapping (genomic/network layouts) | Uses biological structure for positioning | Biologically meaningful | Requires prior knowledge | Pathway-based analysis |
| DeepInsight / structured encoding | Optimized mapping for CNNs | High predictive performance | Computational complexity | Disease classification |
| Hybrid approaches | Combine multiple techniques | Improved robustness | Increased complexity | Multi-modal integration |
This comparative framework highlights that no single method is universally optimal—the choice depends on the biological question, data type, and downstream application.
Deep Learning Models for Image-Based Disease Classification
Once multi-omics data is transformed into images, a wide range of deep learning architectures can be applied. These models excel at identifying complex, hierarchical, and nonlinear patterns, making them particularly suited for biomedical data.
1. Convolutional Neural Networks (CNNs)
CNNs are the dominant architecture for image-based multi-omics analysis.
Key capabilities:
- Detect local spatial patterns (e.g., co-expressed genes)
- Learn hierarchical features across layers
- Scale effectively with large datasets
Clinical applications:
- Cancer subtype classification
- Tumor vs normal tissue prediction
- Molecular phenotype identification
CNN-based models often achieve state-of-the-art performance, frequently outperforming classical machine learning approaches.
2. Transfer Learning
Transfer learning leverages pre-trained models (e.g., ImageNet-trained CNNs) and adapts them to multi-omics images.
Advantages:
- Reduces need for large datasets
- Improves model generalization
- Accelerates training
This is particularly valuable in genomics, where sample sizes are often limited.
3. Autoencoders and Variational Autoencoders (VAEs)
Autoencoders are used for:
- Dimensionality reduction
- Learning latent representations
- Noise reduction
VAEs extend this by learning probabilistic feature distributions, which can improve robustness and interpretability.
4. Generative Adversarial Networks (GANs)
GANs are increasingly used to:
- Generate synthetic multi-omics data
- Address data scarcity
- Improve model training
They are particularly useful in rare diseases where datasets are small.
5. Graph Neural Networks (GNNs) in Hybrid Models
Although not purely image-based, GNNs are often combined with image representations:
- Model biological networks (e.g., gene interactions)
- Complement spatial image features
- Improve interpretability
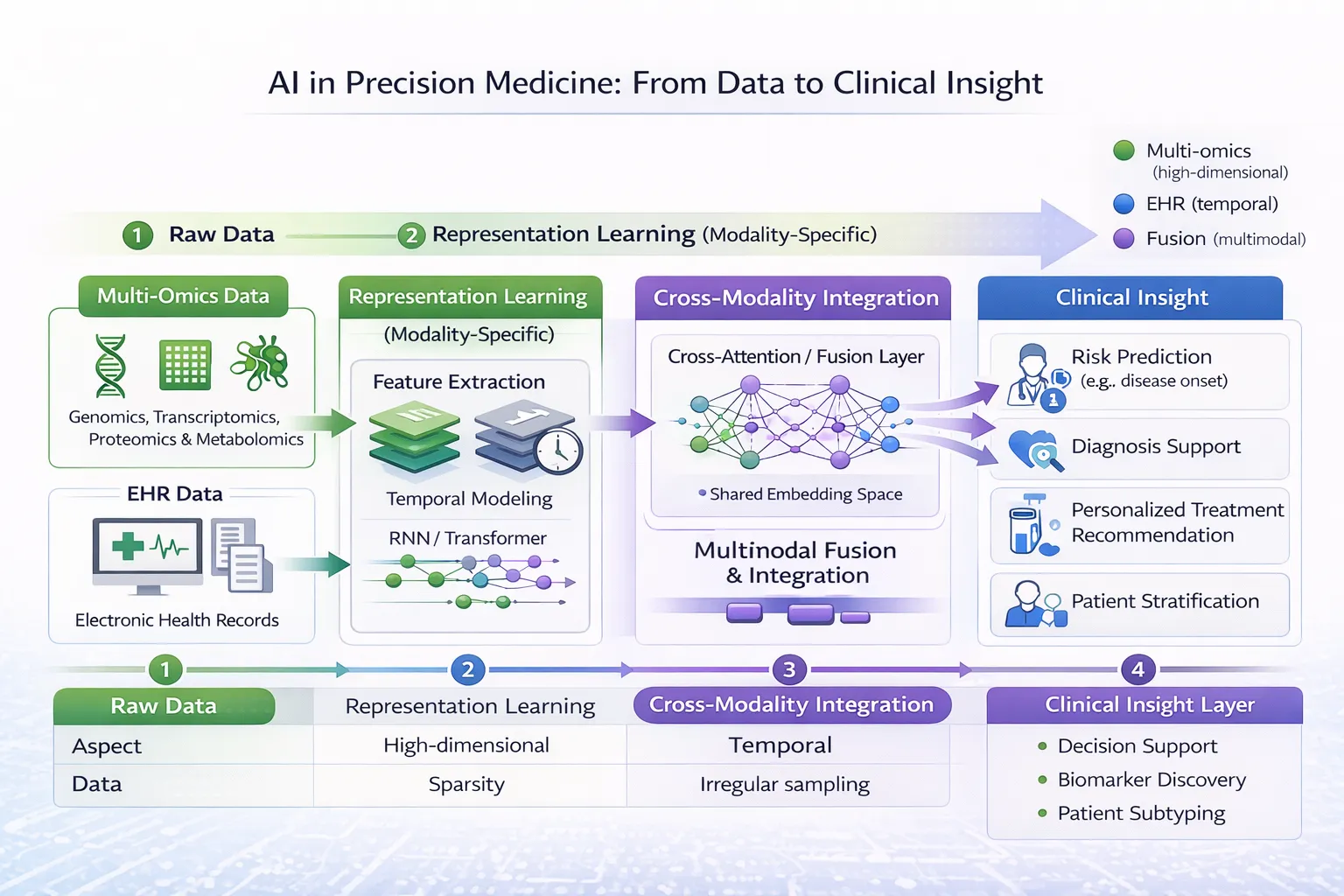
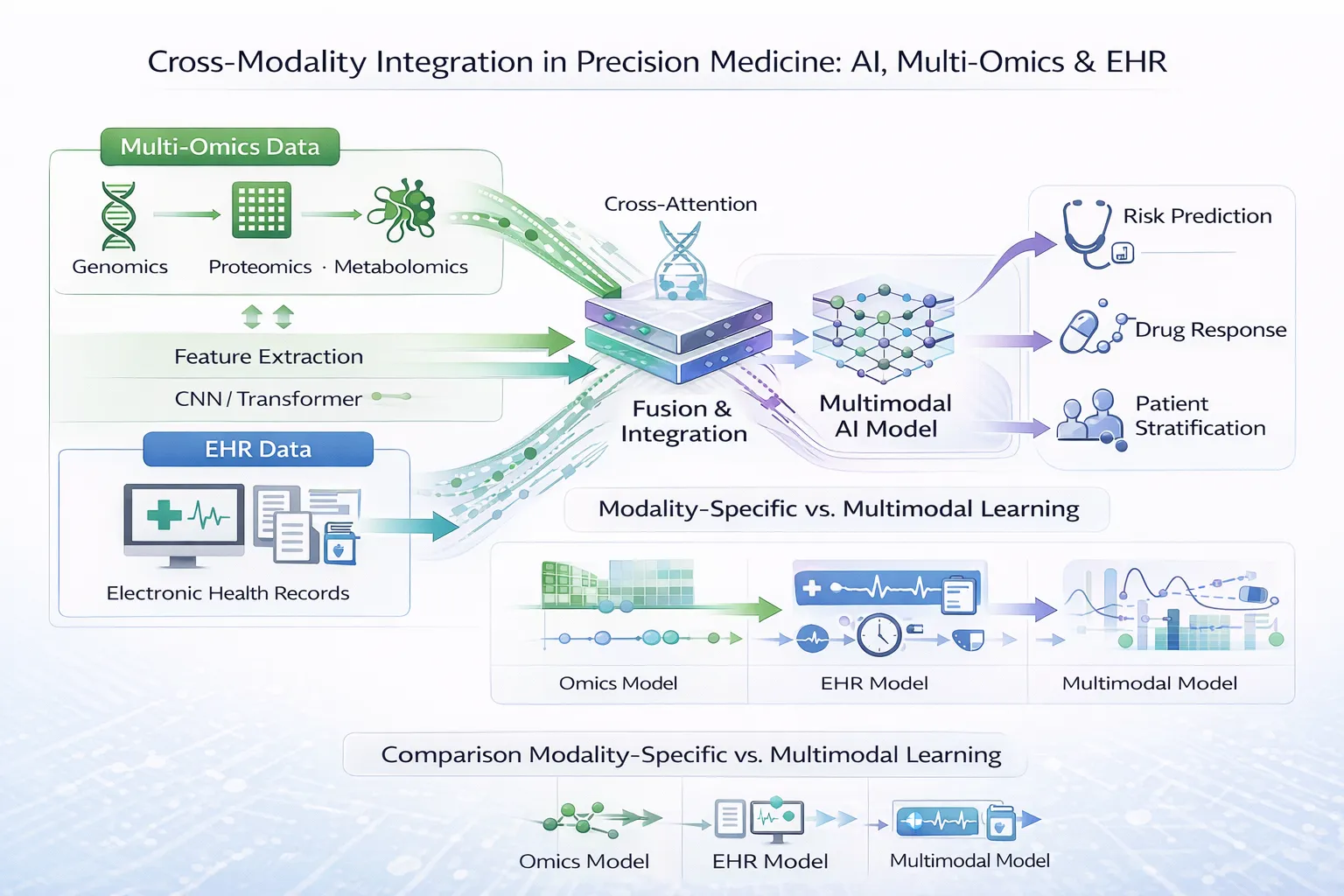
6. Multimodal Deep Learning Architectures
Modern approaches integrate:
- Multi-omics images
- Clinical data (EMRs)
- Imaging data (radiology, pathology)
These models enable holistic disease modeling, reflecting real-world clinical complexity.
Performance and Clinical Impact
Across studies:
- Classification accuracies often range from 75% to >95%
- Improved detection of subtle disease subtypes
- Enhanced ability to predict outcomes and treatment response
These advances are driving the integration of AI into precision medicine workflows.
Applications in Biomedical Research and Medicine
Disease Classification
Image-based multi-omics models can distinguish between disease subtypes with high accuracy, particularly in cancer.
Biomarker Discovery
Spatial representations enable the identification of key molecular signatures associated with disease.
Survival Prediction
Deep learning models can integrate multi-omics images to predict patient outcomes.
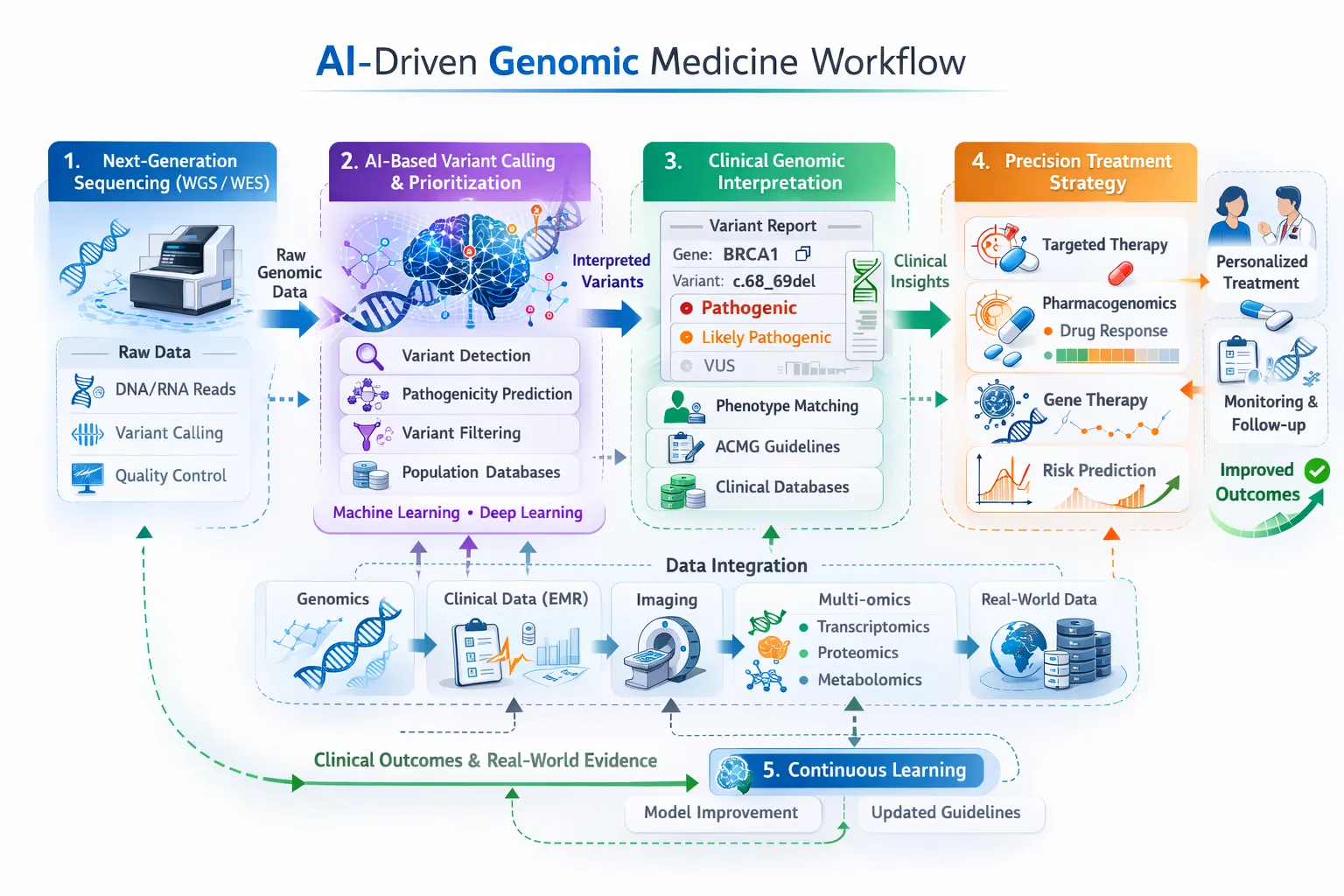
Precision Medicine
By combining multiple biological layers, clinicians can:
- Stratify patients more effectively
- Select targeted therapies
- Improve treatment outcomes
Advantages of Image-Based Multi-Omics Approaches
- Enhanced feature extraction from complex datasets
- Ability to model nonlinear relationships
- Compatibility with state-of-the-art AI models
- Improved performance compared to traditional methods
Importantly, these methods allow researchers to leverage decades of advances in computer vision, accelerating progress in biomedical data analysis.
Challenges and Limitations
Despite their promise, several challenges remain:
Overfitting
Deep learning models may perform well on training data but fail to generalize.
Interpretability
Image-based models can be difficult to interpret biologically.
Data Heterogeneity
Different omics platforms produce inconsistent data types.
Small Sample Sizes
Many multi-omics datasets have limited patient numbers, affecting model robustness.
Standardization
There is currently no universally accepted framework for image transformation.
Addressing these challenges will be essential for clinical adoption. ([ScienceDirect][1])
Image-Based vs Alternative Multi-Omics Integration Methods
| Approach | Strengths | Limitations |
|---|---|---|
| Image-based transformation | Strong pattern recognition, CNN compatibility | Requires preprocessing, interpretability challenges |
| Graph-based models | Capture biological relationships | Depend on prior knowledge |
| Concatenation methods | Simple integration | Limited scalability |
| Model-based integration | Flexible | Computationally intensive |
Image-based approaches are particularly effective when dealing with high-dimensional and sparse datasets, where spatial representation enhances learning. ([ScienceDirect][1])
Future Directions
The field of image-based multi-omics is evolving rapidly, with several key trends emerging:
- Integration with AI-driven clinical decision systems
- Development of interpretable deep learning models
- Use of larger, more diverse datasets
- Combination with real-world clinical data (EMRs)
- Expansion into radiomics and digital pathology ([PubMed][3])
These advances will further bridge the gap between computational biology and clinical practice.
Conclusion
Transforming multi-omics data into images represents a significant innovation in biomedical data science. By enabling the application of deep learning techniques, this approach unlocks new opportunities for disease understanding, diagnosis, and precision medicine.
While challenges remain, continued advances in methodology, data integration, and interpretability are likely to position image-based multi-omics as a cornerstone of next-generation healthcare analytics.