Introduction
The integration of genomics with longitudinal clinical data represents a fundamental shift toward truly personalized medicine. While Electronic Health Records (EHRs) encode temporal patient trajectories—including diagnoses, procedures, medications, and laboratory measurements—they lack direct representation of inherited disease risk. Conversely, genomic data, particularly polygenic risk scores (PRS), capture cumulative genetic susceptibility but lack temporal and environmental context.
The study “Integrating Genomics into Multimodal EHR Foundation Models” 🔗 proposes a unified modeling framework that embeds genomic information directly into EHR-based foundation models. By treating genomics as a native modality rather than an auxiliary feature, the approach enables joint learning of genotype–phenotype relationships across time.
Data: Multimodal Cohort Construction and Representation
1. Data Source: All of Us Research Program
The model is trained on data derived from the All of Us (AoU) Research Program, a large-scale, longitudinal cohort designed to support precision medicine research. The dataset integrates:
Structured EHR data
- Diagnoses (ICD codes)
- Procedures
- Medications
- Laboratory measurements
Genomic data
- Genome-wide genotyping
- Derived polygenic risk scores (PRS) for multiple traits
The dataset is notable for:
- Scale: Hundreds of thousands of participants
- Diversity: Inclusion of underrepresented populations
- Longitudinality: Multi-year patient timelines
- Multimodality: Linked clinical and genomic records
2. Representation of EHR Data
EHR data are represented as temporal sequences of discrete clinical events, where:
- Each event is tokenized (e.g., diagnosis code, medication code)
- Events are ordered chronologically
- Temporal structure is preserved implicitly through sequence position
This formulation allows the use of sequence modeling architectures (transformers) similar to those used in natural language processing.
3. Representation of Genomic Data
Genomic information is incorporated via polygenic risk scores, which are:
- Scalar or vector representations summarizing genetic risk
- Derived from genome-wide association studies (GWAS)
- Computed across thousands of genetic variants
Key design choice:
PRS are embedded into the model input space, enabling joint optimization with EHR representations.
This contrasts with traditional pipelines where PRS are appended post hoc to prediction models.
4. Multimodal Alignment
A central challenge addressed in the paper is aligning heterogeneous modalities:
| Modality | Structure | Challenge |
|---|---|---|
| EHR | Temporal sequence | Variable length, irregular timing |
| Genomics | Static vector | No temporal dimension |
The model resolves this by:
- Encoding each modality separately
- Projecting both into a shared latent representation space
- Learning cross-modal interactions during pretraining
Methodology: Multimodal Foundation Model Architecture
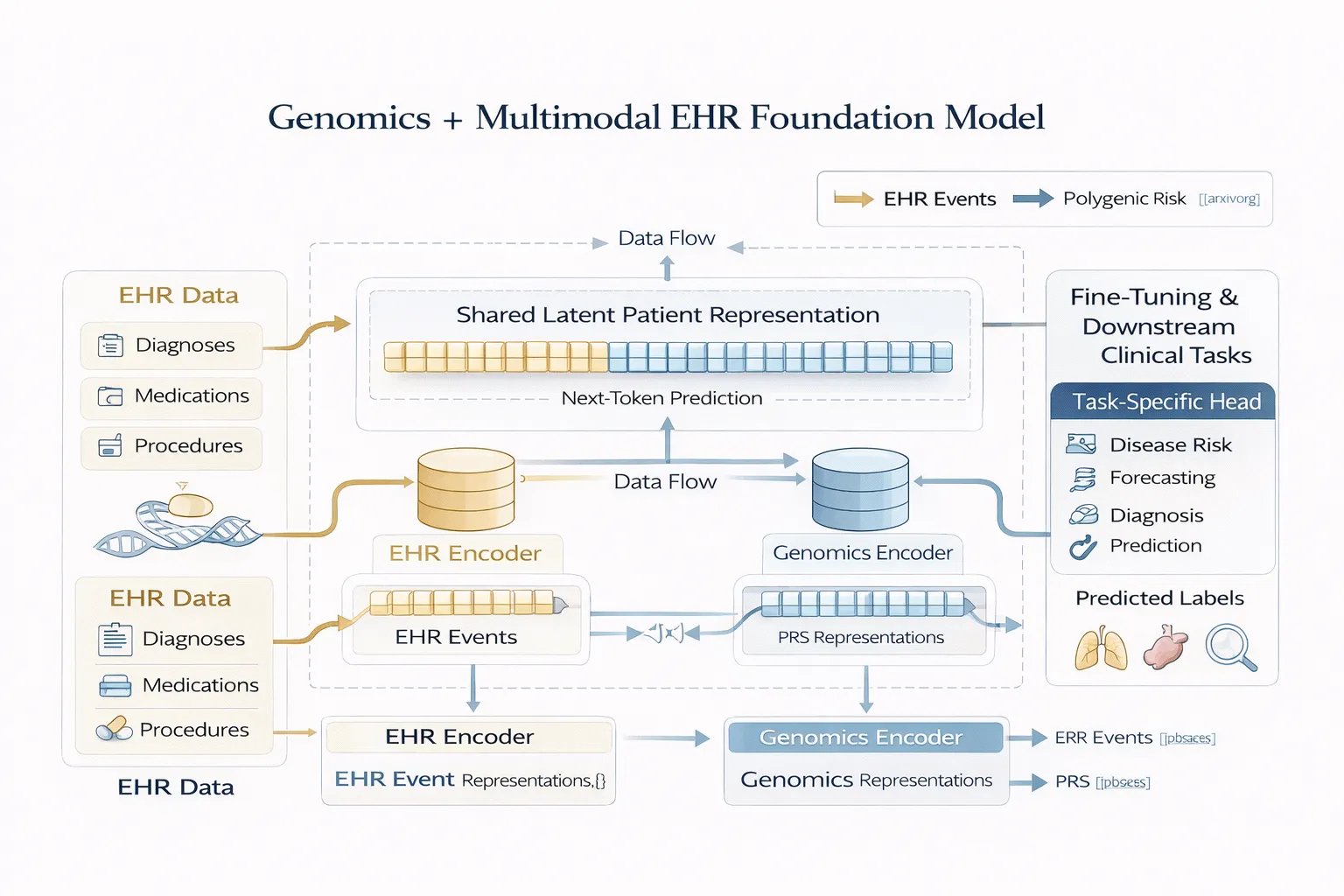
1. Overall Architecture
The proposed model extends transformer-based EHR foundation models with a dual-encoder multimodal design:
Components:
- EHR Encoder
- Transformer-based sequence model
- Processes longitudinal clinical tokens
- Captures temporal dependencies
- Genomics Encoder
- Feedforward embedding network
- Maps PRS into dense representations
- Fusion Mechanism
- Combines modality-specific embeddings
- Produces a shared latent patient representation
- Task Head(s)
- Fine-tuned for downstream prediction tasks
2. Pretraining Objective
The model is trained using a self-supervised generative objective, specifically:
Next-Token Prediction
- Given a sequence of clinical events, predict the next event
- Analogous to language modeling (e.g., GPT-style training)
Formally:
P(x_t | x_1, x_2, ..., x_{t-1}, \text{PRS})Where:
- (x_t) = clinical event at time (t)
- PRS = genomic input
This objective enables the model to learn:
- Disease progression patterns
- Temporal dependencies in clinical data
- Interactions between genetic risk and clinical events
3. Multimodal Fusion Strategy
Fusion occurs at the representation level, not decision level:
- PRS embeddings are integrated into the model input or intermediate layers
- Attention mechanisms enable cross-modal interactions
- The model learns how genomic risk influences clinical trajectories
This approach allows:
- Early fusion (joint representation learning)
- Better generalization compared to late fusion approaches
4. Fine-Tuning and Downstream Tasks
After pretraining, the model is fine-tuned for clinical prediction tasks such as:
- Disease onset prediction
- Risk stratification
- Clinical outcome forecasting
Fine-tuning involves:
- Adding task-specific heads
- Training on labeled datasets
- Leveraging pretrained representations
Key Results and Findings
1. Performance Gains from Genomic Integration
The study demonstrates that:
- Multimodal models (EHR + PRS) outperform EHR-only models
- Improvements are observed across multiple disease domains
- Genomics contributes non-redundant predictive signal
2. Early Risk Prediction
Incorporating PRS enables:
- Identification of high-risk individuals before clinical manifestation
- Improved long-term prediction horizons
3. Representation Learning Benefits
The shared latent space captures:
- Cross-modal dependencies
- Latent disease trajectories
- Interactions between inherited risk and observed outcomes
Discussion: Technical Insights and Implications
1. Genomics as a First-Class Modality
A key conceptual shift in this work is treating genomics as a first-class input modality, rather than:
- A covariate
- A post hoc feature
This allows the model to learn joint distributions over clinical and genetic data, improving both:
- Predictive performance
- Biological relevance
2. Foundation Model Paradigm in Healthcare
The study reinforces the applicability of foundation models in healthcare:
- Large-scale pretraining enables generalizable representations
- Transfer learning reduces need for labeled data
- Multimodal learning captures complex biological systems
3. Modeling Genotype–Phenotype Relationships
By integrating PRS into temporal models, the framework enables:
- Implicit modeling of genotype–phenotype relationships
- Context-aware interpretation of genetic risk
- Dynamic risk estimation over time
4. Data Efficiency and Scalability
The use of self-supervised learning allows:
- Efficient utilization of large unlabeled datasets
- Scalability to additional modalities
Limitations
The paper identifies several limitations inherent to the current approach:
- Dependence on PRS Representation
- PRS compress genomic information into summary statistics
- May not capture fine-grained variant-level effects
- Modality Imbalance
- EHR data is high-dimensional and temporal
- Genomics is static and lower dimensional
- This asymmetry can affect representation learning
- Model Interpretability
- The integration of modalities within deep architectures reduces transparency
- Understanding how genomic features influence predictions remains challenging
Future Directions
The study outlines several directions for future work:
- Richer Genomic Representations Moving beyond PRS to include raw genomic features by incorporating variant-level or sequence-level data
- Improved Multimodal Fusion Mechanisms: Developing more sophisticated architectures for cross-modal interaction
- Expansion to Additional Modalities: Integrating imaging, proteomics, or wearable data
- Enhanced Interpretability Methods: Developing tools to better understand multimodal model outputs
Conclusion
Integrating genomics into multimodal EHR foundation models represents a significant advancement in AI-driven healthcare. By embedding polygenic risk scores directly into the learning process, the proposed framework enables more accurate and biologically meaningful predictions.
This work demonstrates the feasibility and value of joint multimodal representation learning, paving the way toward fully integrated precision medicine systems.
Explore related insights
- Multi-omics Applications in Human Diseases
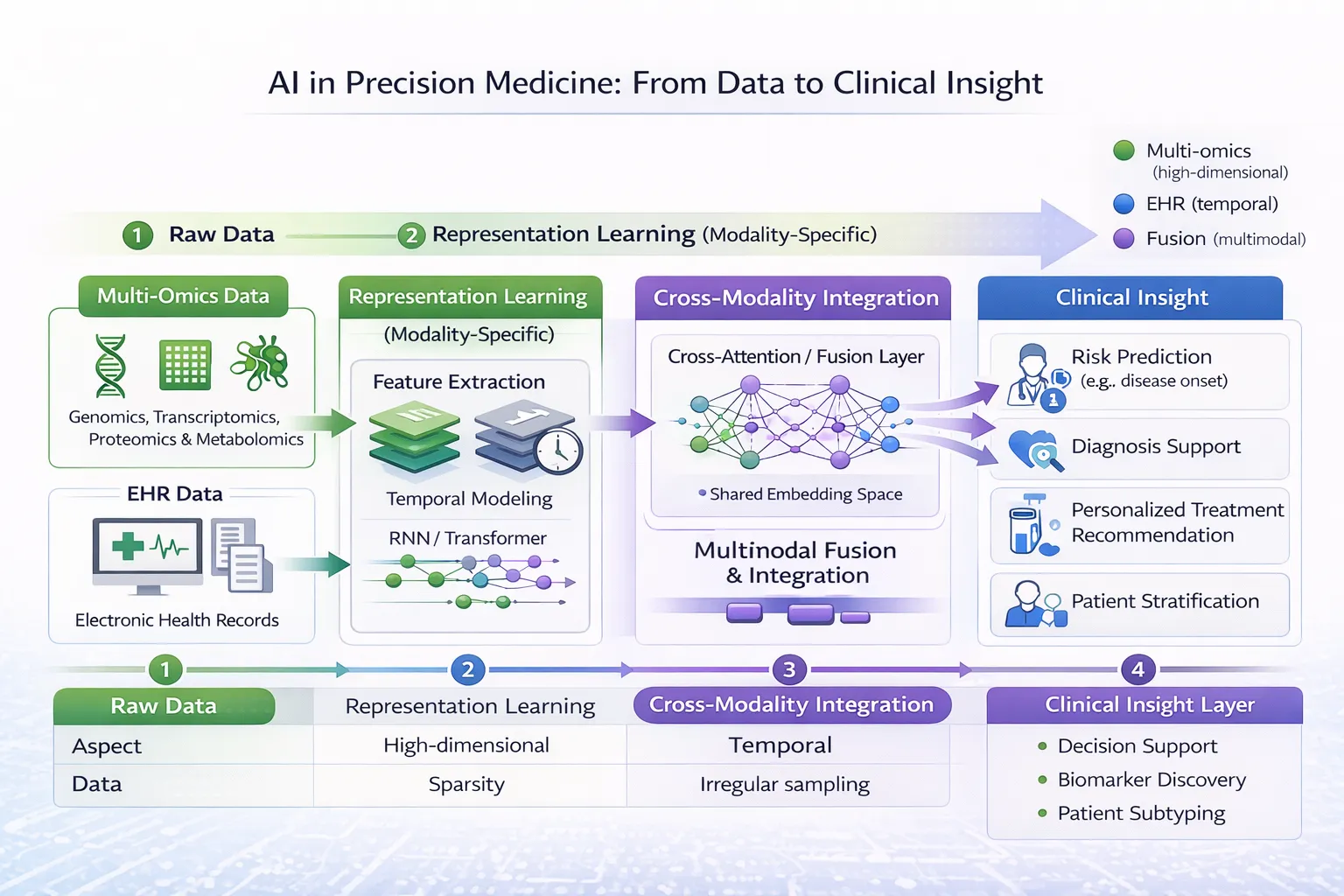
- AI for Multi-Omics and EHR Data: Modality-Specific Learning
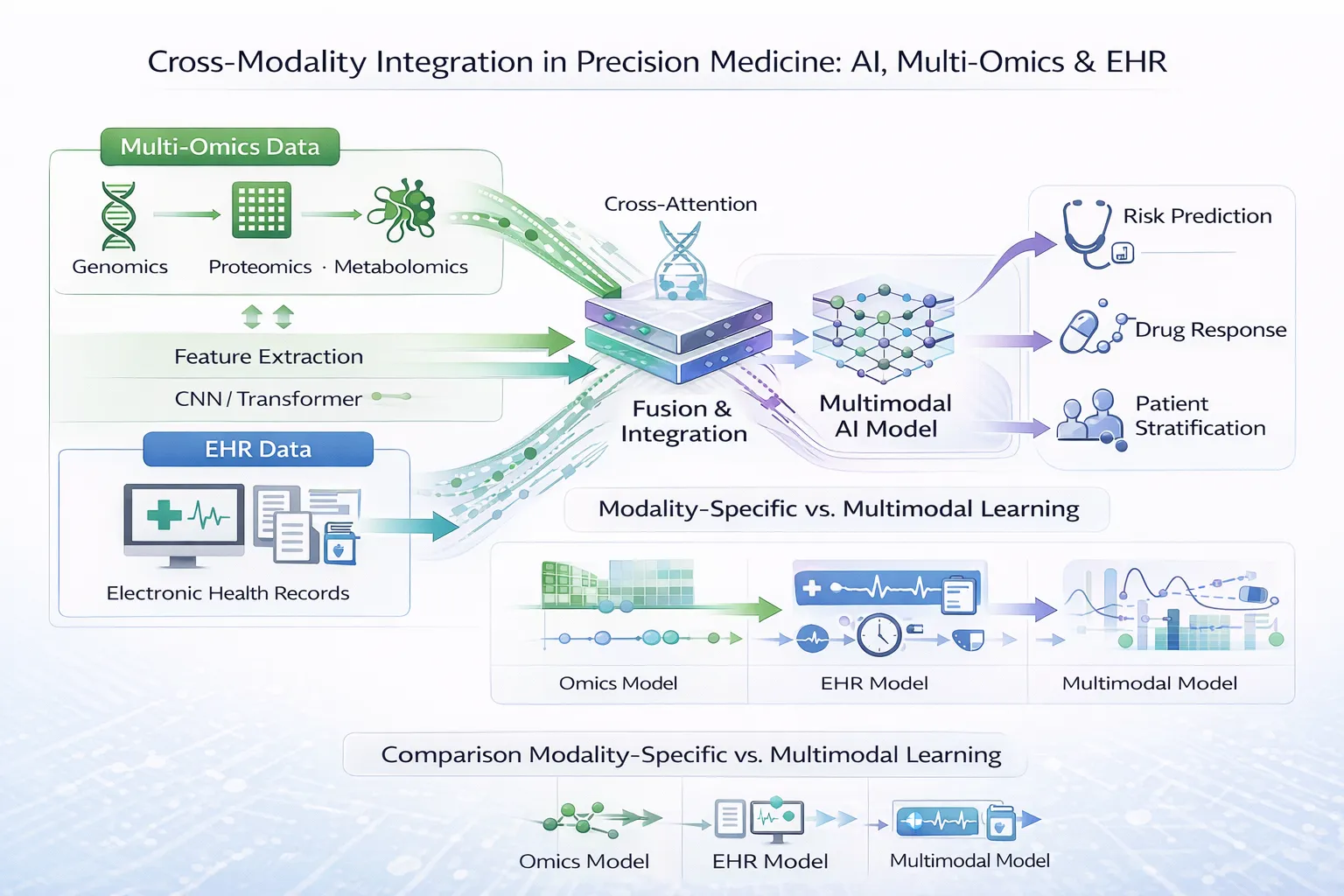
- Cross-Modality Integration in Precision Medicine
- Deep Learning for Multi-Omics Data Integration