Introduction
Modern healthcare generates vast and complex datasets spanning molecular biology and clinical practice. However, transforming this data into actionable clinical insight remains a major challenge. Multi-omics datasets capture the biological foundations of disease, while electronic health records (EHRs) provide longitudinal, real-world patient information.
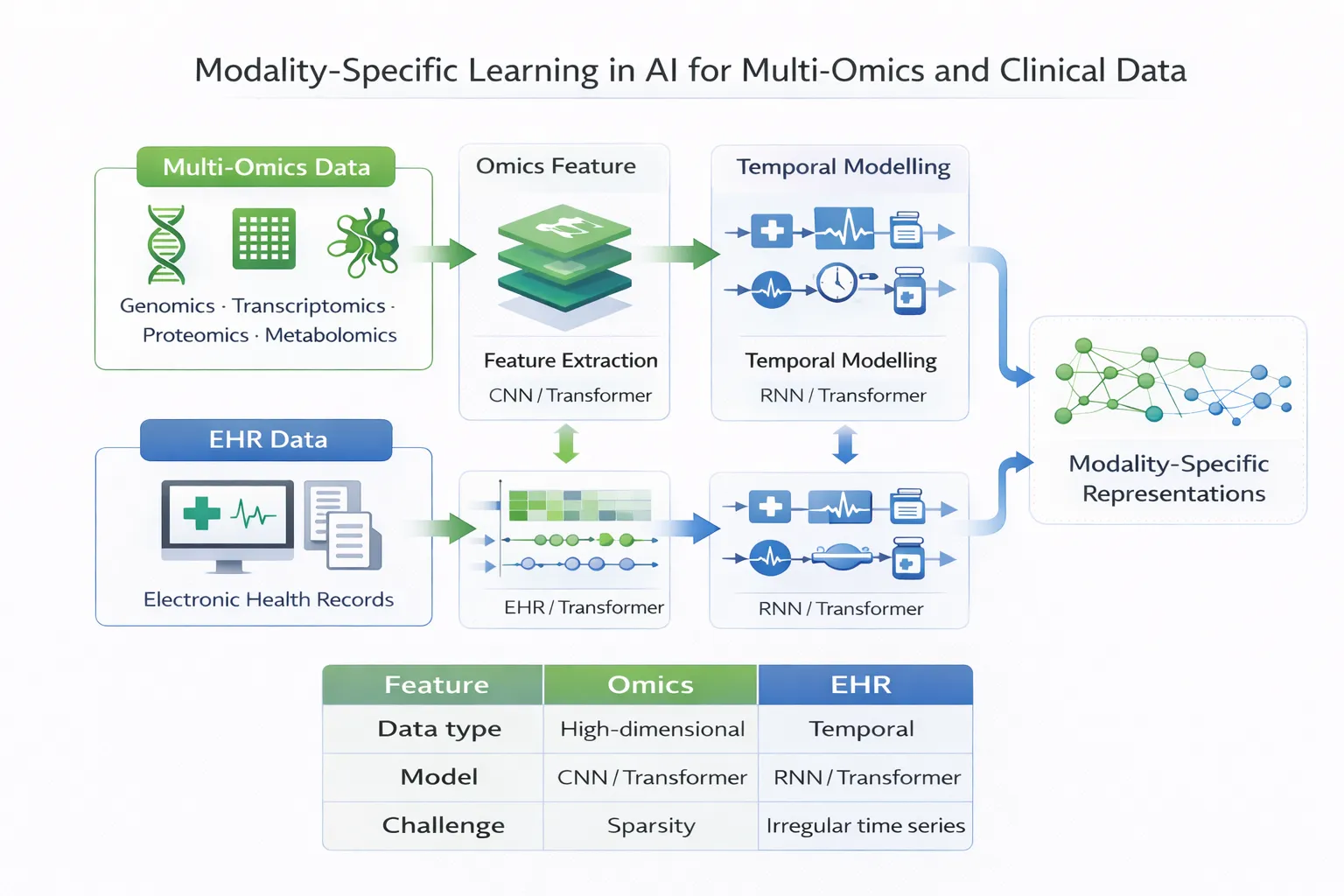
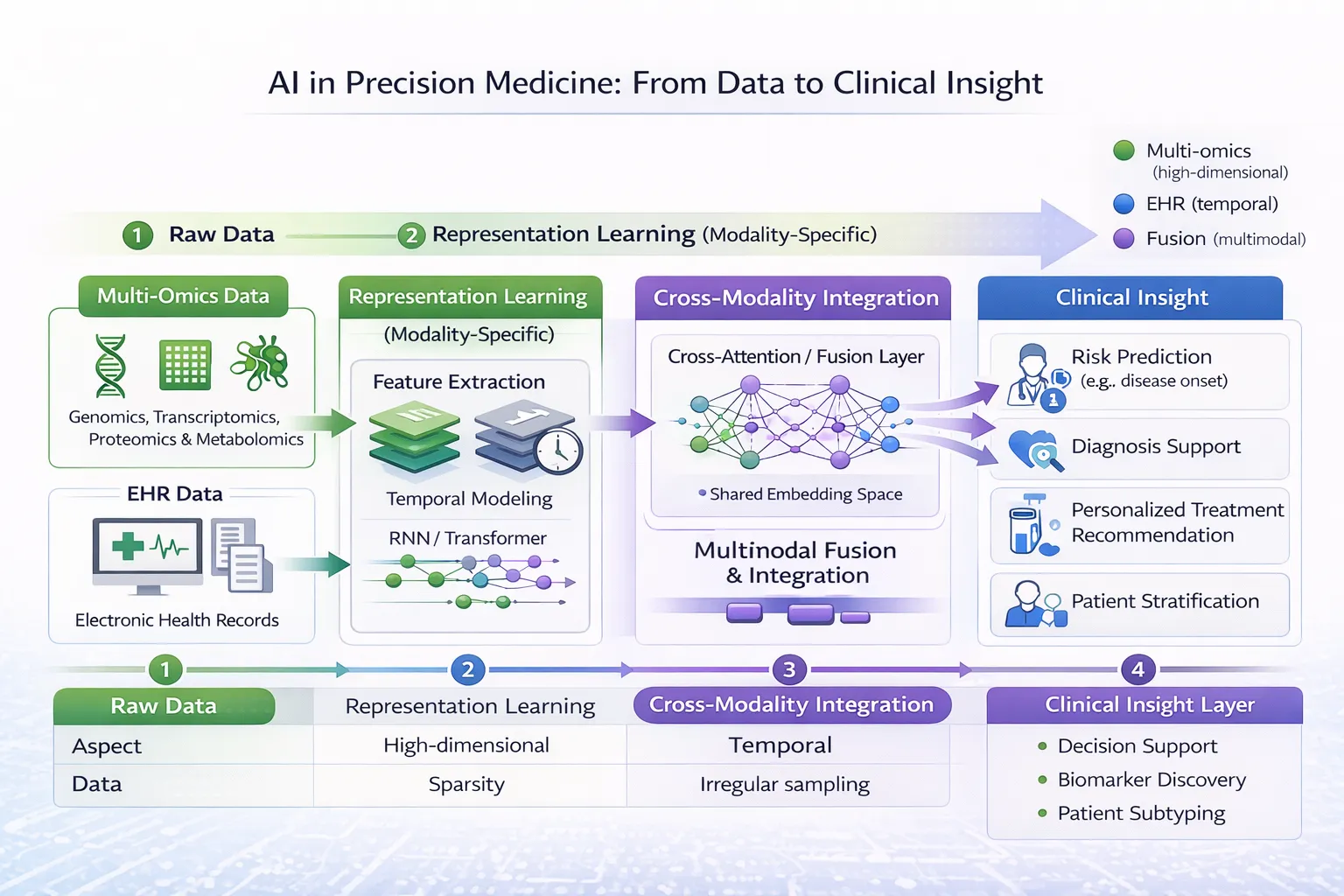
Artificial intelligence (AI) is increasingly enabling modality-specific learning—tailored analytical approaches designed to extract meaningful patterns from each data type independently. These methods form a critical foundation for precision medicine, ensuring that each modality is properly understood before integration.
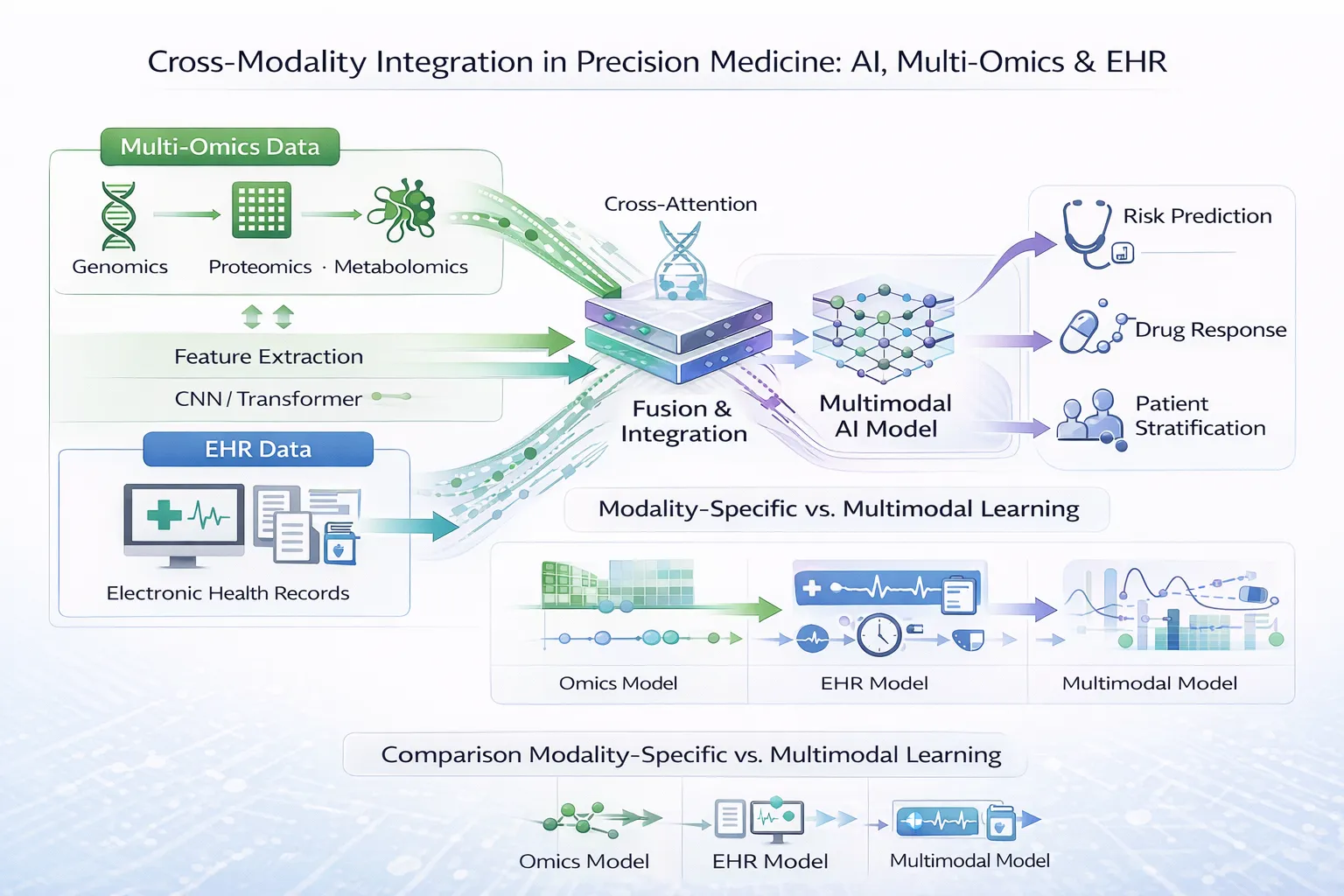
This is the first part of our AI in Precision Medicine: From Data to Clinical Insight series. Check out Part 2: Cross-Modality Integration in Precision Medicine too!
Key Takeaways
- Multi-omics data requires dimensionality reduction and representation learning to manage high dimensionality
- EHR data requires temporal and multimodal modelling to capture patient trajectories
- Modality-specific learning improves robustness and interpretability prior to integration
- Molecular and clinical data provide complementary insights for precision medicine
Section 1: Modality-Specific Learning for Multi-Omics Data
Understanding Multi-Omics Data
Multi-omics data encompasses multiple layers of biological information, including:
- Genomics (DNA variation)
- Transcriptomics (gene expression)
- Proteomics (protein abundance)
- Metabolomics (metabolic activity)
These datasets are characterised by:
- Extremely high dimensionality (thousands to millions of variables)
- Relatively small sample sizes
- Complex biological interdependencies
This imbalance leads to the well-known curse of dimensionality, where traditional statistical approaches struggle to generalise effectively.
Key Challenges in Multi-Omics Learning
High Dimensionality and Limited Samples
The number of molecular features far exceeds the number of patients, increasing overfitting risk and limiting model robustness.
Biological Heterogeneity
Different omics layers represent distinct biological processes, each with unique distributions and scales.
Noise and Batch Effects
Experimental variability introduces noise, while missing data is common across platforms.
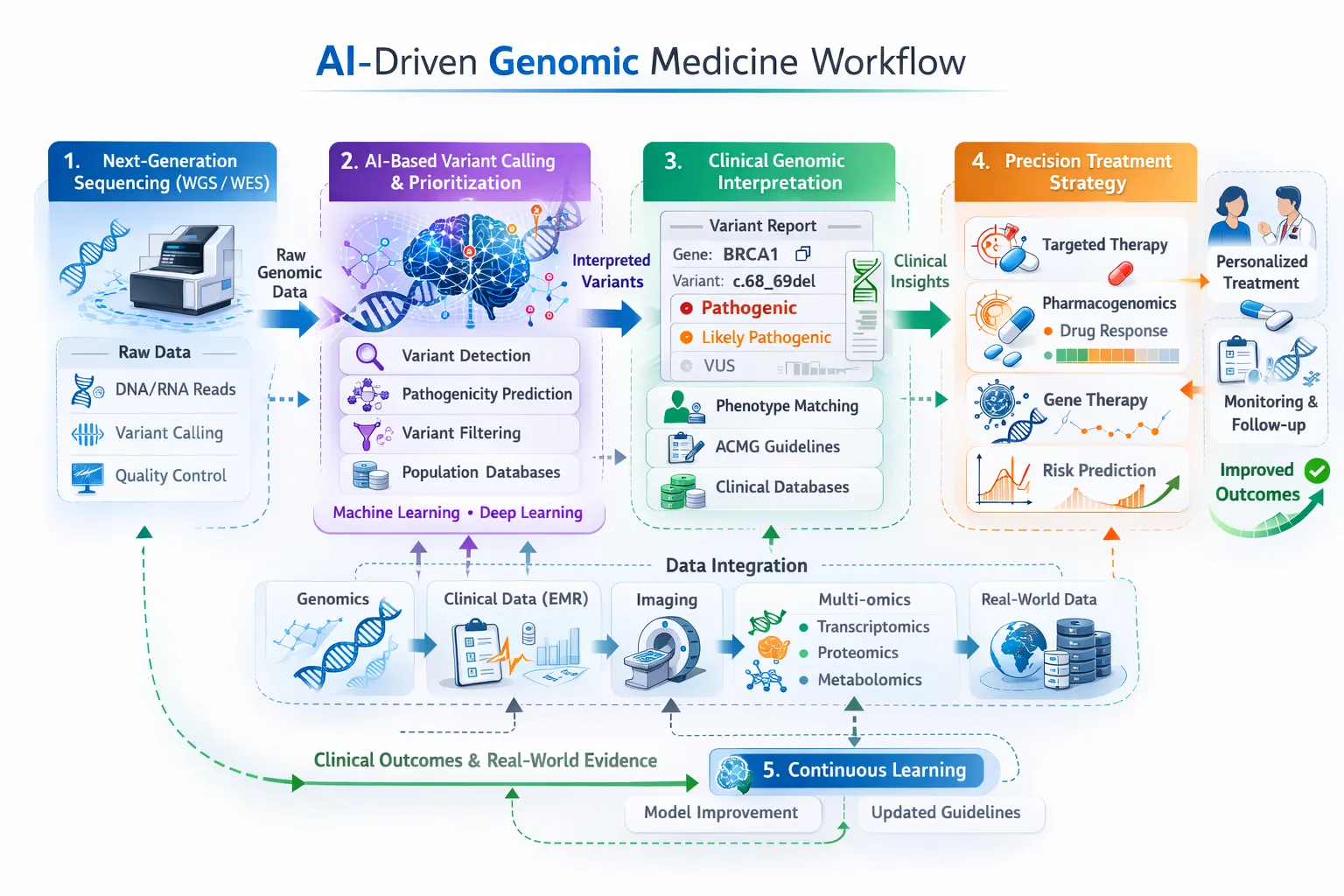
AI Approaches for Multi-Omics Data
1. Dimensionality Reduction
Techniques such as:
- Principal Component Analysis (PCA)
- Autoencoders
- Variational Autoencoders (VAEs)
are widely used to:
- Compress high-dimensional data
- Extract latent biological structure
- Improve downstream prediction tasks
These methods reduce noise while preserving key biological signals.
2. Representation Learning
Deep learning models enable the learning of latent embeddings that capture complex molecular relationships.
- Autoencoders map multi-omics data into compact representations
- Graph Neural Networks (GNNs) model biological networks and interactions
- Feature selection methods (e.g., LASSO, mutual information) identify relevant biomarkers
Such approaches have demonstrated improved performance in cancer classification and survival prediction¹.
3. Task-Specific Modelling
Once features are extracted, models are applied to:
- Disease classification
- Risk stratification
- Biomarker discovery
Clinical example: Multi-omics integration has been used to identify breast cancer subtypes, enabling more precise treatment selection and improved patient outcomes².
Applications in Precision Medicine
Modality-specific learning for multi-omics enables:
- Identification of disease subtypes
- Prediction of therapeutic response
- Discovery of novel drug targets
These insights provide the molecular foundation of personalised medicine.
Section 2: Modality-Specific Learning for Electronic Health Records (EHR)
What Makes EHR Data Unique?
EHR data differs fundamentally from omics data, comprising:
- Structured data (lab results, diagnoses, medications)
- Unstructured data (clinical notes, reports)
- Temporal data (longitudinal patient histories)
EHRs provide a real-world, time-resolved view of patient health, making them indispensable for clinical decision-making.
Key Challenges in EHR Learning
Data Heterogeneity
EHRs combine numerical, categorical, and textual data, requiring flexible modelling approaches.
Sparsity and Missingness
Clinical data is often incomplete and non-randomly missing, reflecting real-world clinical workflows.
Noise and Bias
Coding inconsistencies, documentation errors, and institutional biases can affect model reliability.
AI Approaches for EHR Data
1. Structured Data Modelling
Traditional machine learning methods (e.g., logistic regression, random forests) are used for:
- Risk prediction
- Disease classification
However, these models are limited in capturing temporal dynamics.
2. Temporal and Sequential Models
Deep learning approaches such as:
- Recurrent Neural Networks (RNNs)
- Long Short-Term Memory (LSTM) networks
- Transformer-based models (e.g., BEHRT, Med-BERT)
are designed to model patient trajectories over time, capturing:
- Disease progression
- Treatment response
- Temporal dependencies
3. Multimodal Representation Learning
Modern approaches treat EHRs as multi-view datasets, integrating:
- Clinical measurements
- Text data via natural language processing (NLP)
- Imaging summaries
These models learn latent representations that reflect underlying disease states³.
Clinical Applications of EHR Learning
EHR-based AI models support:
Early disease detection Example: Predicting sepsis hours before clinical onset
Hospital readmission prediction Identifying high-risk patients for targeted intervention
Mortality risk estimation Supporting ICU decision-making
Clinical decision support systems Enhancing physician decision-making at the point of care
EHR-driven AI is increasingly embedded in healthcare systems to improve outcomes and operational efficiency⁴.
Comparing Multi-Omics and EHR Learning
| Feature | Multi-Omics Data | EHR Data |
|---|---|---|
| Data type | Molecular | Clinical |
| Structure | High-dimensional | Heterogeneous |
| Key challenge | Dimensionality | Temporal complexity |
| Typical models | Autoencoders, GNNs | RNNs, Transformers |
| Clinical role | Mechanistic insight | Real-world patient context |
Why Modality-Specific Learning Matters
While both data types are essential, they require fundamentally different analytical approaches.
- Multi-omics learning uncovers biological mechanisms
- EHR learning captures clinical trajectories
Careful modality-specific modelling ensures:
- Reduced noise and bias
- Robust feature extraction
- Improved downstream integration
This step is critical before applying cross-modality learning, where these datasets are combined.
Clinical Relevance
For clinicians and healthcare systems, modality-specific learning enables:
- Earlier and more accurate diagnosis
- Improved risk stratification
- Personalised treatment planning
- Enhanced clinical decision support
By bridging molecular biology and patient care, these approaches are transforming how medicine is practised.
Conclusion
Modality-specific learning represents a foundational pillar of precision medicine. By tailoring AI models to the unique characteristics of multi-omics and EHR data, researchers can extract meaningful insights from each domain independently.
As AI continues to evolve, these approaches will play an increasingly critical role in translating complex biomedical data into actionable clinical knowledge, ultimately improving patient outcomes.
In the Part 2 of this series, we explore how these modalities are combined through cross-modality integration learning, along with the challenges and opportunities this presents.