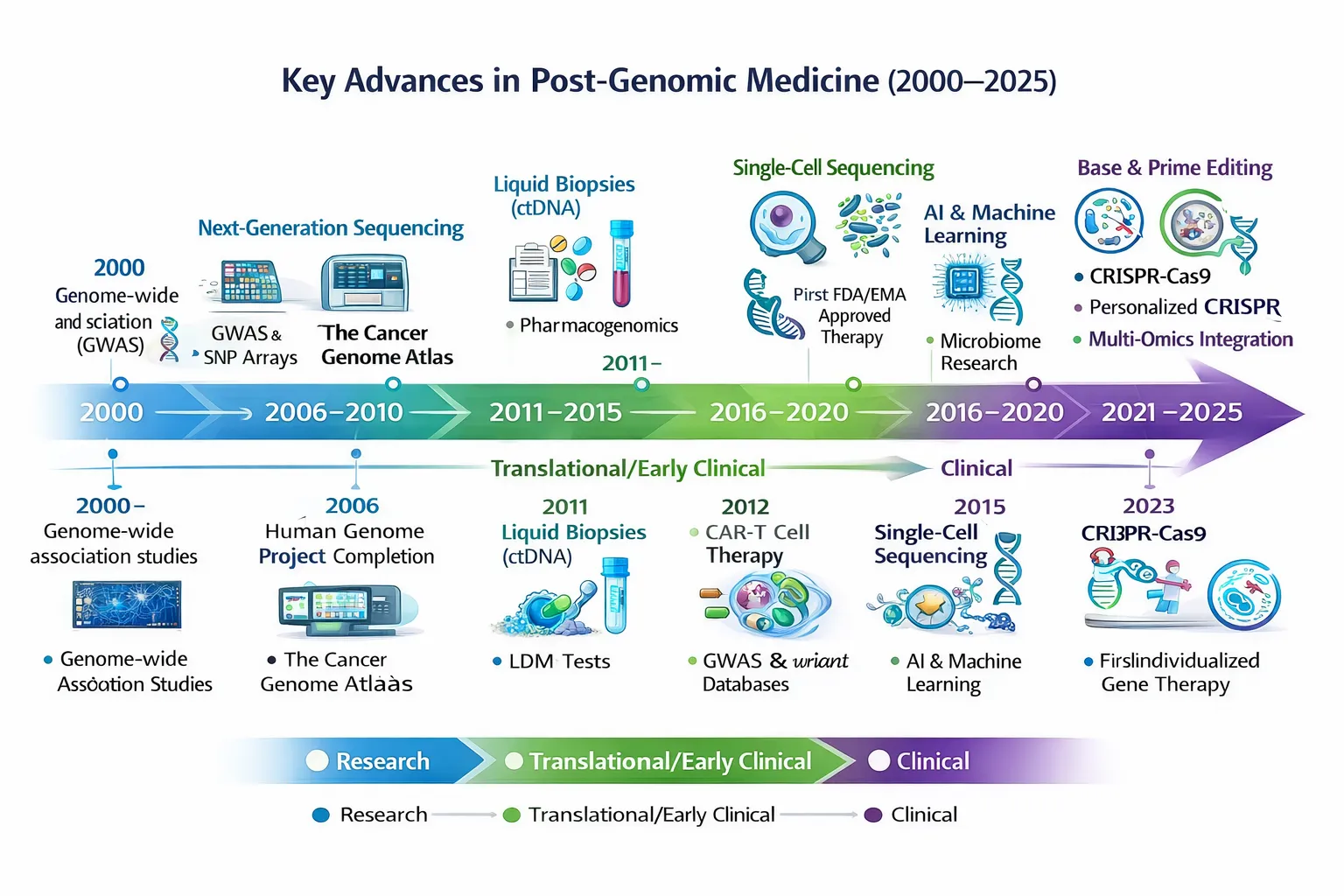
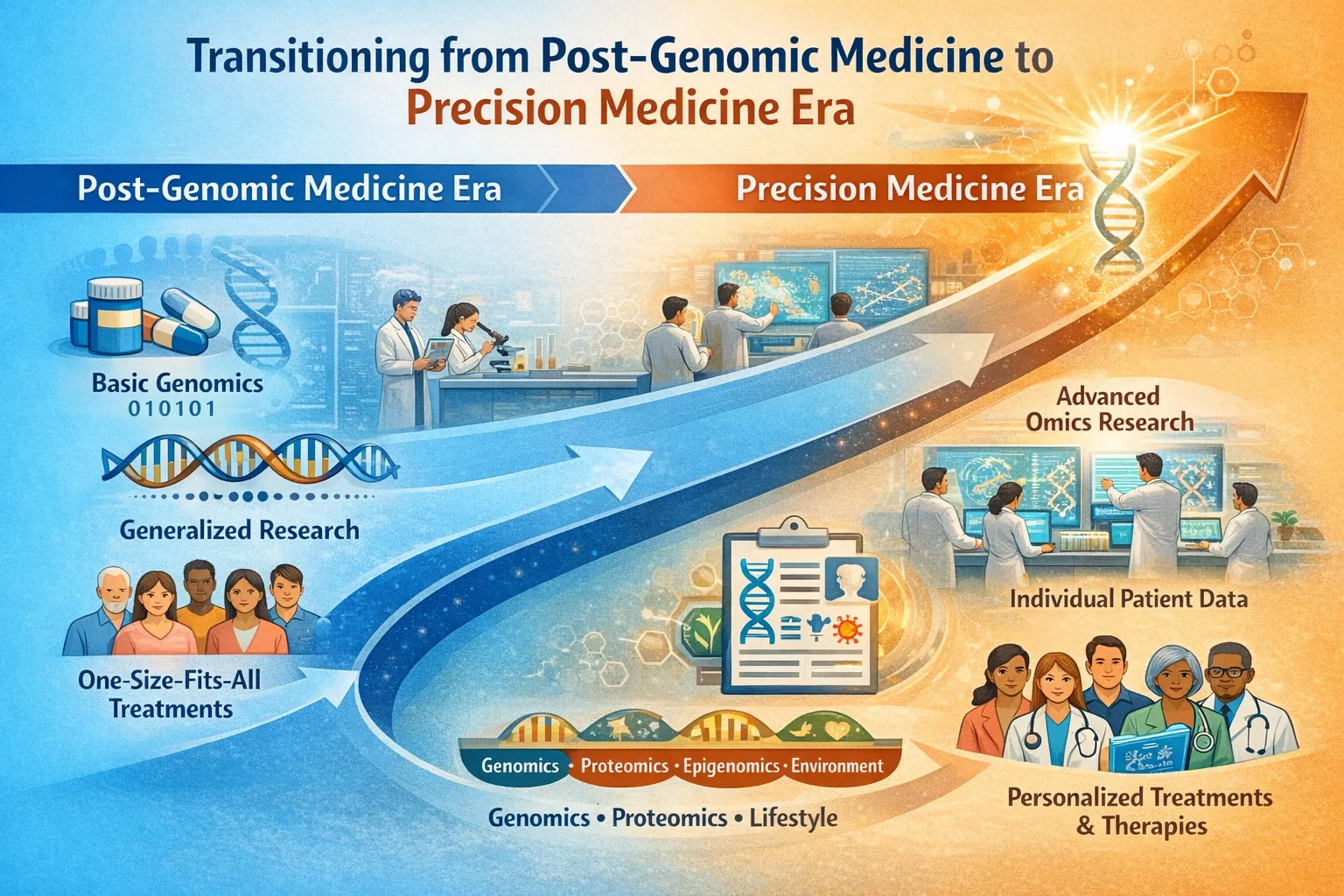
Introduction
Over the last 30+ years, we’ve witnessed a transformation in how medicine understands and treats disease. The shift didn’t happen overnight — it unfolded as biology went from decoding what DNA is to deciphering how DNA and biology behave in context. This transition defines post‑genomic medicine: a discipline grounded in genome data but extending into multi‑omics, systems biology, and personalized care.
In this post, we’ll unpack what post‑genomic medicine means, how it differs from earlier approaches, and why it matters — with real examples and published evidence.
A Timeline of Human Genomics: From Pre‑Genome to Post‑Genome
The completion of the Human Genome Project marked a transformative moment in biomedical science, promising a future where decoding DNA would unlock the secrets of disease. While genomics has delivered important insights, it quickly became clear that DNA sequence alone cannot fully explain human health and disease.
We have now entered the post-genomic era—a paradigm shift that moves beyond static genetic information toward a dynamic, systems-level understanding of biology. Post-genomic medicine integrates multiple layers of biological data—collectively known as multi-omics—alongside clinical and environmental information to enable more precise, predictive, and personalized healthcare.
| Era | Milestone | What Changed |
|---|---|---|
| Pre‑Genomic Era | 1953 – Discovery of DNA structure | Watson & Crick reveal the double helix, laying the foundation for molecular genetics. |
| 1980s – Single‑gene studies | Diseases like cystic fibrosis and Duchenne muscular dystrophy are mapped to individual genes. | |
| Genomic Era | 1990–2003 – Human Genome Project (HGP) | First complete human genome sequence provides a foundational reference. |
| 2005–2010 – High‑throughput genotyping & sequencing | Genome‑wide scans identify many disease‑associated variants. | |
| Post‑Genomic Era | 2010s – Multi‑omics & systems biology | Integration of genomics with transcriptomics, proteomics, metabolomics. |
| 2020s – Human pangenome & precision medicine delivery | Diverse reference genomes and clinical translation initiatives proliferate. |
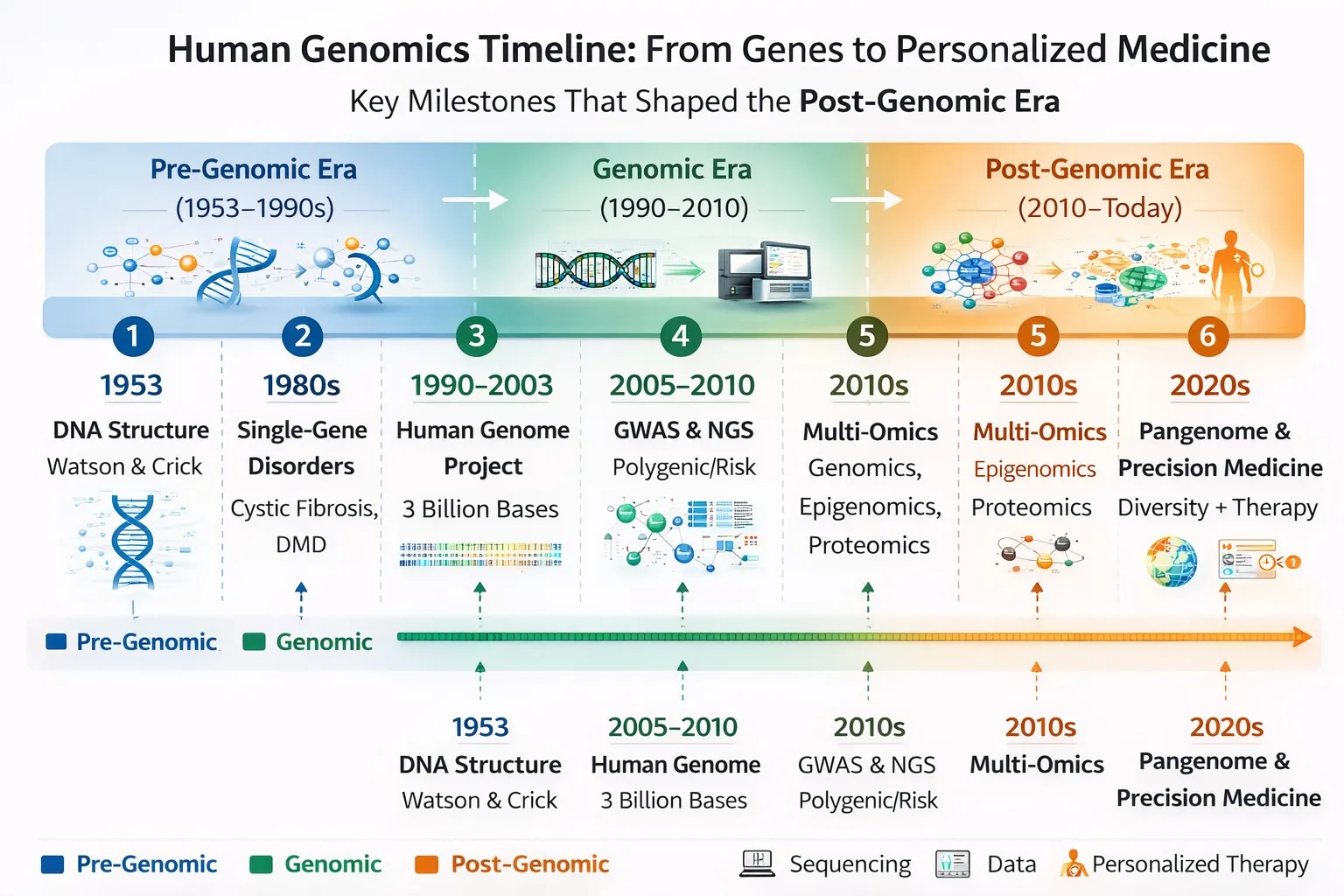
Key Differences: Pre‑Genomic vs Post‑Genomic
A key distinction in modern medicine is the transition from a gene-centric view to a systems-based approach.
| Feature | Pre‑Genomic | Post‑Genomic |
|---|---|---|
| Approach | Gene-centric | Systems-integrated |
| Predictive power | Limited | Probabilistic & multi‑factorial |
| Diagnostics | Symptom → test | Sequencing → interpretation → tailored care |
| Data | Single genes | Multi‑omics + clinical + environmental |
This shift reflects a fundamental insight: genes do not act in isolation. Instead, health outcomes arise from complex interactions between genes, molecular processes, environment, and time.
What “Post‑Genomic” Really Means
In the pre‑genomic and genomic eras, research focused on genes in isolation: find a mutation, link it to a disease. But in real biology, and especially in medicine, disease emerges from interactions:
- Between genes and regulatory networks
- Between genome and environment
- Between immune system and cells
- Across tissues, time, and life stages
This shift is why the term post‑genomic medicine exists.
📌 Sensemaking, not just sequencing, is now central.
What is Multi-Omics?
Post-genomic medicine is built on the integration of multiple biological layers, each providing unique insights into human biology. This extends beyond genes into the expression of everything downstream to genes.
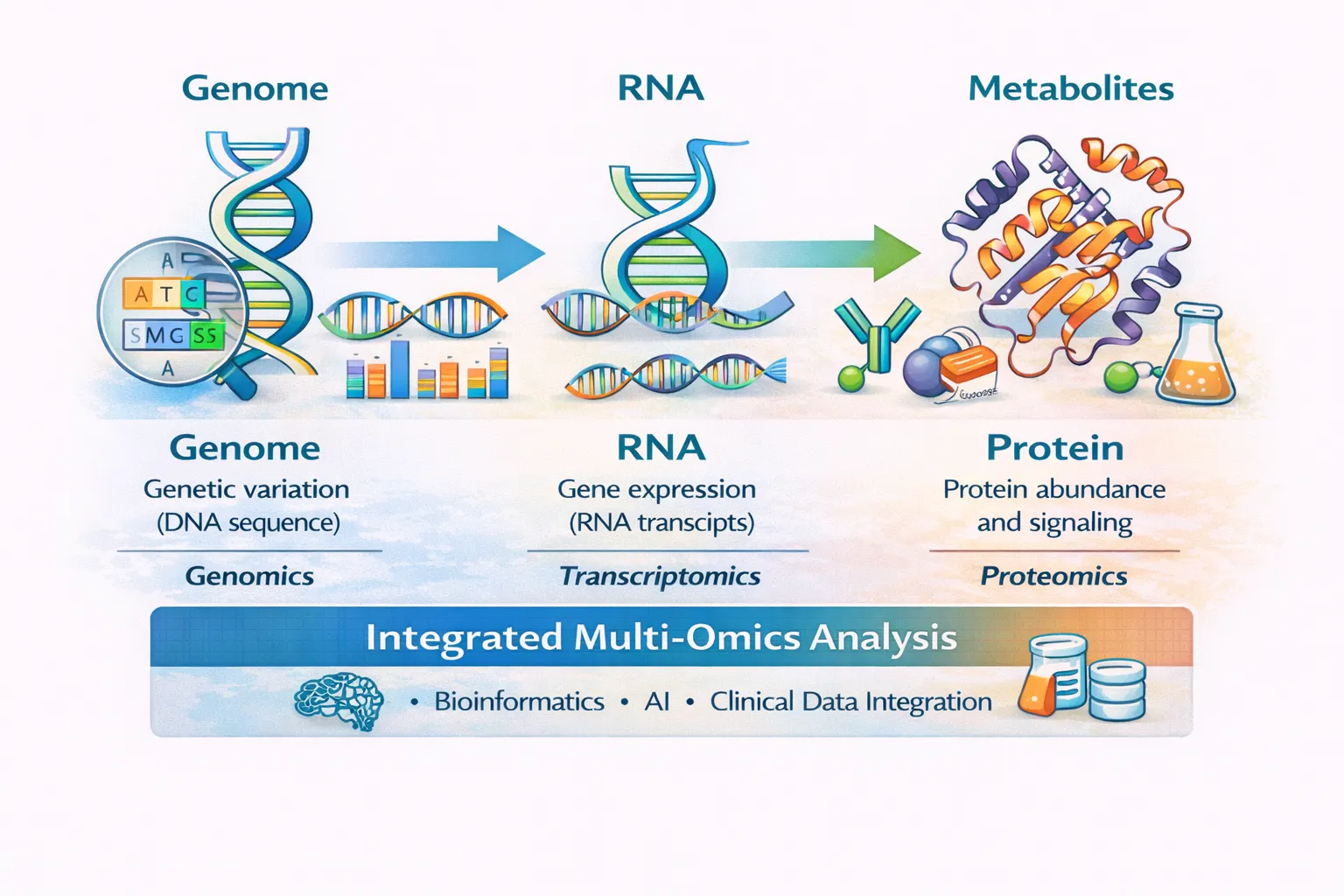
Genomics
Genomics examines the DNA sequence—the foundational blueprint of life. It identifies inherited and acquired variants that may influence disease risk.
Example: Mutations in BRCA1 and BRCA2 increase the risk of breast and ovarian cancer.
Transcriptomics
Transcriptomics studies RNA expression, revealing which genes are actively being transcribed in a given cell or tissue.
Example: Tumor gene expression profiling helps classify cancer subtypes and guide treatment decisions.
Proteomics
Proteomics focuses on proteins—the functional molecules that carry out most biological processes.
Example: HER2 protein overexpression in breast cancer guides the use of targeted therapies such as trastuzumab.
Metabolomics
Metabolomics analyzes small molecules (metabolites) that reflect real-time cellular activity and physiological state.
Example: Altered glucose and lipid metabolites are key indicators in diabetes and metabolic disorders.
In other articles, we will cover how these layers of Multi-omics together with Artificial Intelligence play a crucial role in the future of post genomic medicine.
Examples That Define the Post‑Genomic Era
Below are real, examples showing post‑genomic medicine in action:
1. Beyond Single Genes: Multi‑Gene Risk and GWAS
Early genome‑wide association studies (GWAS) revealed that common diseases like type 2 diabetes and coronary artery disease are influenced by hundreds of genetic variants, each with a small effect.
- Publication example: Diabetes GWAS identifying dozens of loci associated with disease risk (Nature Genetics, 2018)
- Clinical implication: No single mutation predicts disease — risk must be modeled across the genome.
What it means to the patient: DNA isn’t destiny — it contributes probabilistically.
What it means to the clinician: Genetic risk scores enhance stratification but must be used with caution due to effect sizes and ancestry bias.
2. Next‑Generation Sequencing (NGS): From Research to Clinic
NGS reduced sequencing cost and time dramatically, enabling whole‑exome and whole‑genome sequencing in clinical diagnostics.
- Identifies pathogenic variants in undiagnosed diseases with diagnostic yields of ~30–40% in rare disease cohorts (JAMA, 2019)
- Tumor sequencing reveals actionable targets for therapy (Cancer Cell, 2017) .
What it means to the patient: A child with unexplained seizures receives a diagnosis after whole‑genome sequencing.
What it means to the clinician: NGS guides targeted therapies in oncology — e.g., FLT3 mutations dictating FLT3 inhibitor use in AML.
3. Cancer as a Genomic Disease
The Cancer Genome Atlas (TCGA) and related efforts defined the molecular landscapes of multiple cancers.
- Integrated multi‑omic profiling revealed distinct subtypes of glioblastoma with prognostic relevance (NEJM, 2010) .
- EGFR mutations in non‑small cell lung cancer predict response to EGFR tyrosine kinase inhibitors (NEJM, 2004) .
What it means to the patient: Two cancers that look the same under a microscope can behave very differently at the molecular level.
What it means to the clinician: Genomic profiling is now standard in many tumors to select targeted therapies.
4. Human Pangenome and Equity
Traditional reference genomes were based on a single or limited set of ancestries, biasing variant interpretation. The human pangenome project aims to address this.
- Multi‑ancestry references improve detection of structural variants and reduce diagnostic disparities (Nature, 2023) .
What it means to the patient: Our genomes are diverse, and references must reflect that.
What it means to the clinician: Awareness of ancestry effects improves variant interpretation and reduces false positives/negatives.
Why This Matters for Patients (and Clinicians)
For patients
Post-genomic medicine has direct implications for everyday healthcare:
- Explains why patients respond differently to the same treatment
- Faster diagnoses for rare and complex conditions
- More personalized treatments rather than one‑size‑fits‑all
- Better risk prediction and preventive strategies
For clinicians
- Integration of genomic data into routine care
- Nuanced understanding of variant pathogenicity
- Collaboration with genetics specialists and molecular tumor boards
- Recognition of limitations: VUS, ancestry biases, environmental modifiers
Ultimately, it shifts medicine from a reactive model to a proactive and preventive approach.
Conclusion
Post-genomic medicine represents a fundamental evolution in healthcare. By integrating multi-omics data, clinical information, and advanced computational tools, it moves beyond the limitations of DNA alone to capture the full complexity of human biology.
This shift is not merely technological—it is conceptual. It redefines how we understand disease, how we treat patients, and how we envision the future of medicine.