Introduction
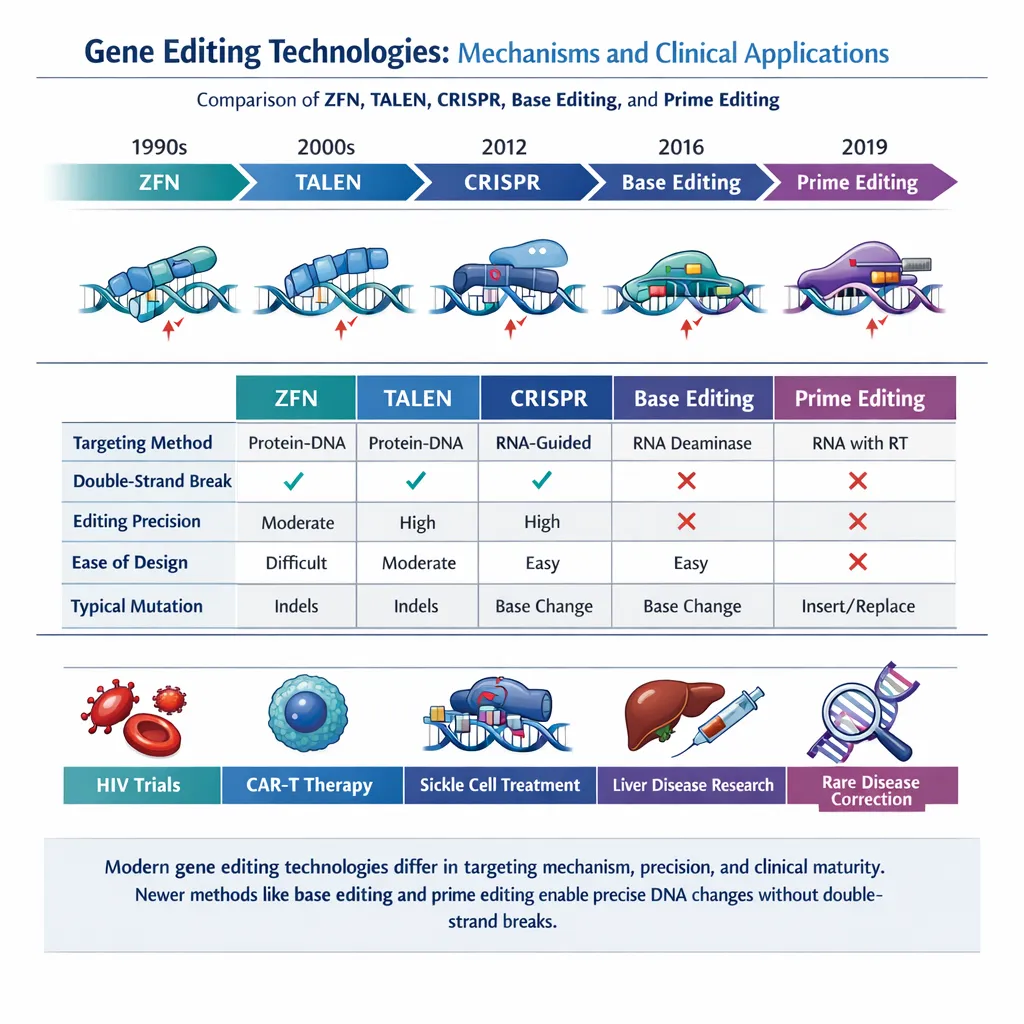
Prime editing (PE) has emerged as a next-generation genome editing technology that overcomes many of the limitations associated with earlier CRISPR-based approaches. By enabling precise, programmable DNA modifications without inducing double-strand breaks (DSBs) or requiring donor templates, PE significantly reduces unintended genomic alterations while expanding the range of editable mutations.
Recent studies have demonstrated its growing potential in therapeutic genome correction, particularly for monogenic diseases, while ongoing innovations are addressing challenges in efficiency, delivery, and specificity.
1. Prime Editing: Basic Mechanism and Specificity
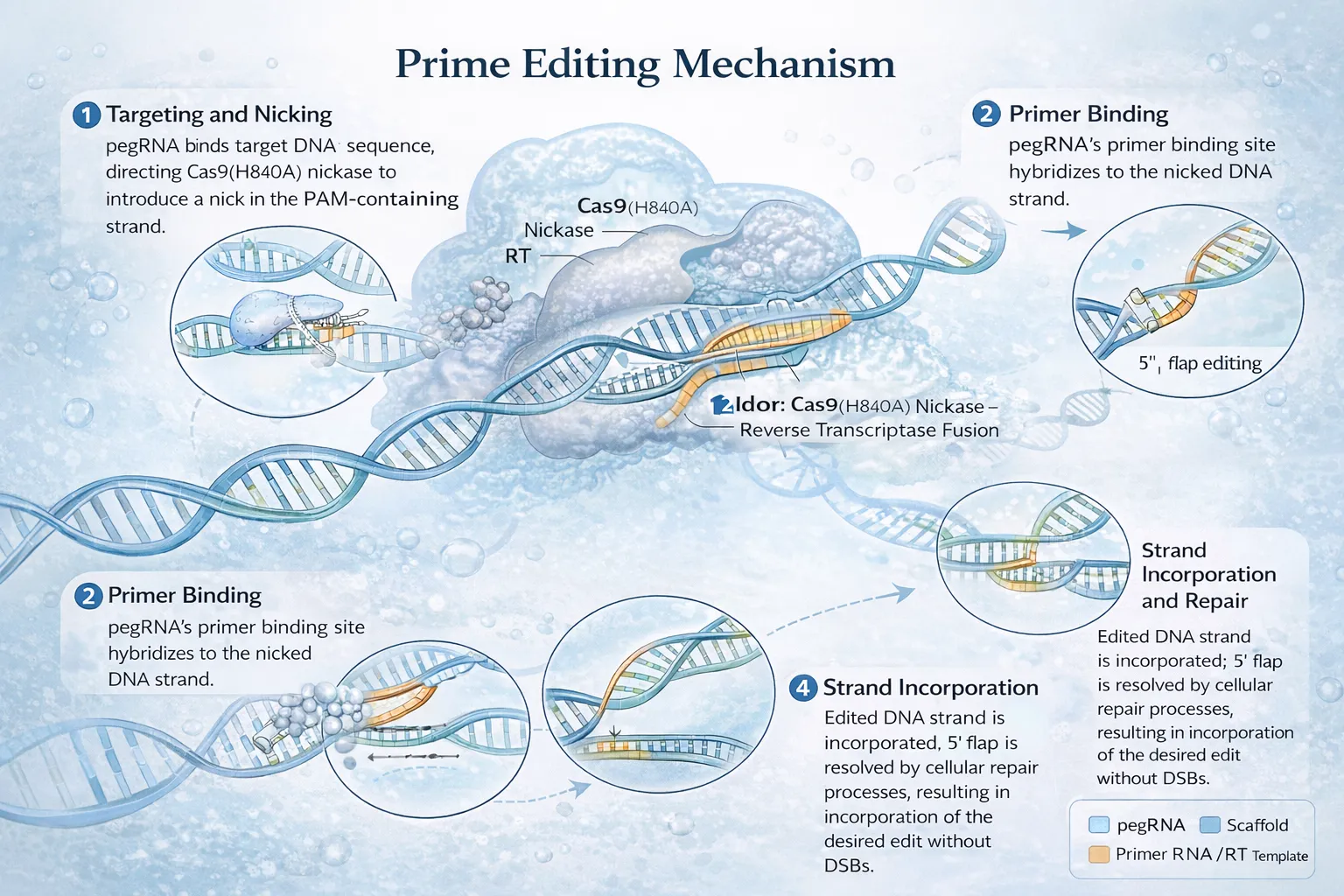
Prime editing relies on a fusion protein composed of Cas9 nickase (nCas9, typically H840A) and an engineered reverse transcriptase (RT), guided by a prime editing guide RNA (pegRNA). This system introduces edits through a reverse transcription-based mechanism rather than relying on endogenous homology-directed repair (HDR).
The editing process proceeds through several coordinated steps:
Target Recognition and Binding The pegRNA directs nCas9 to a specific genomic locus via complementary base pairing and PAM recognition.
Single-Strand Nicking nCas9 introduces a nick in the target DNA strand, generating a 3′ hydroxyl group.
Reverse Transcription Initiation The 3′ end hybridizes with the primer binding site (PBS) in the pegRNA, initiating reverse transcription.
DNA Synthesis The RT extends the nicked strand using the RT template region of the pegRNA, encoding the desired edit.
Strand Resolution and Repair Cellular repair pathways incorporate the edited strand into the genome, replacing the original sequence.
Editing Scope
Prime editing supports:
- All 12 base substitutions
- Small insertions (up to ~40 bp)
- Small deletions
- Combinatorial edits (substitutions + indels)
Specificity and Fidelity
Compared to conventional CRISPR-Cas9:
- No DSBs → reduced genomic instability
- Lower indel formation
- Reduced p53 activation and cytotoxicity
- Enhanced targeting precision through pegRNA design
However, specificity depends on:
- pegRNA design parameters (PBS length, RT template)
- genomic context
- DNA repair pathway activity
2. Mechanisms for Overcoming Current Challenges in Using PE
Despite its versatility, PE faces limitations in editing efficiency, product purity, and scalability, particularly in primary cells and in vivo systems.
2.1 pegRNA Engineering and Optimization
pegRNAs are inherently unstable due to their extended structure.
Key improvements:
- Engineered pegRNAs (epegRNAs) with structured RNA motifs (e.g., pseudoknots)
- Optimized PBS length (8–17 nt) for hybridization efficiency
- Tuned RT template length (10–35 nt) to maximize editing yield
These modifications improve:
- RNA stability
- Target engagement
- Editing efficiency
2.2 Enhanced Prime Editing Systems
PE1 → PE2 → PE3 Evolution
- PE1: Initial system with wild-type RT
- PE2: Improved RT activity → higher efficiency
- PE3: Introduces a second nick on the non-edited strand
- PE3b: Conditional nicking to reduce off-target edits
Trade-offs:
- PE3 increases efficiency but may elevate indel rates
- PE3b improves fidelity but may reduce efficiency
2.3 Modulation of DNA Repair Pathways
DNA repair pathways significantly influence editing outcomes.
Strategies:
- Inhibition of mismatch repair (MMR) increases editing efficiency
- Biasing repair toward flap resolution pathways
- Use of small molecules to modulate repair activity
However, long-term suppression of repair pathways raises safety concerns.
2.4 Protein Engineering and Cas Variants
- Development of Cas variants with relaxed PAM requirements expands targetable regions
- Engineering more efficient RT enzymes improves editing kinetics
2.5 Minimizing Byproducts
Undesired edits include:
- Partial edits
- Indels
- Unintended substitutions
Approaches to reduce these:
- Improved pegRNA design algorithms
- Controlled nicking strategies
- Computational prediction of editing outcomes
3. Developing PE Delivery Strategies
Efficient delivery of PE components remains one of the primary bottlenecks for clinical translation.
3.1 Viral Delivery Systems
Adeno-Associated Virus (AAV)
- High transduction efficiency
- Tissue specificity
Limitations:
- Packaging size (~4.7 kb) insufficient for full PE system
- Requires dual-AAV strategies
3.2 Dual-AAV and Split Systems
In this approach Prime Editing components are split across two vectors. These components are then reassembled in vivo via:
- Inteins
- Protein complementation
Challenges:
- Reduced efficiency
- Increased complexity
3.3 Non-Viral Delivery Approaches
Non viral delivery approaches are also being explored, of which two approaches are prominent.
Lipid Nanoparticles (LNPs)
- Deliver mRNA encoding PE components
- Reduced immunogenicity
- Transient expression → improved safety
Electroporation
- Used for ex vivo editing of stem cells and immune cells
3.4 Emerging Delivery Technologies
- Virus-like particles (VLPs) for protein delivery
- Nanocarriers and polymer systems
- Targeted delivery systems for specific tissues
Delivery remains a critical determinant of:
- Therapeutic efficacy
- Tissue specificity
- Safety
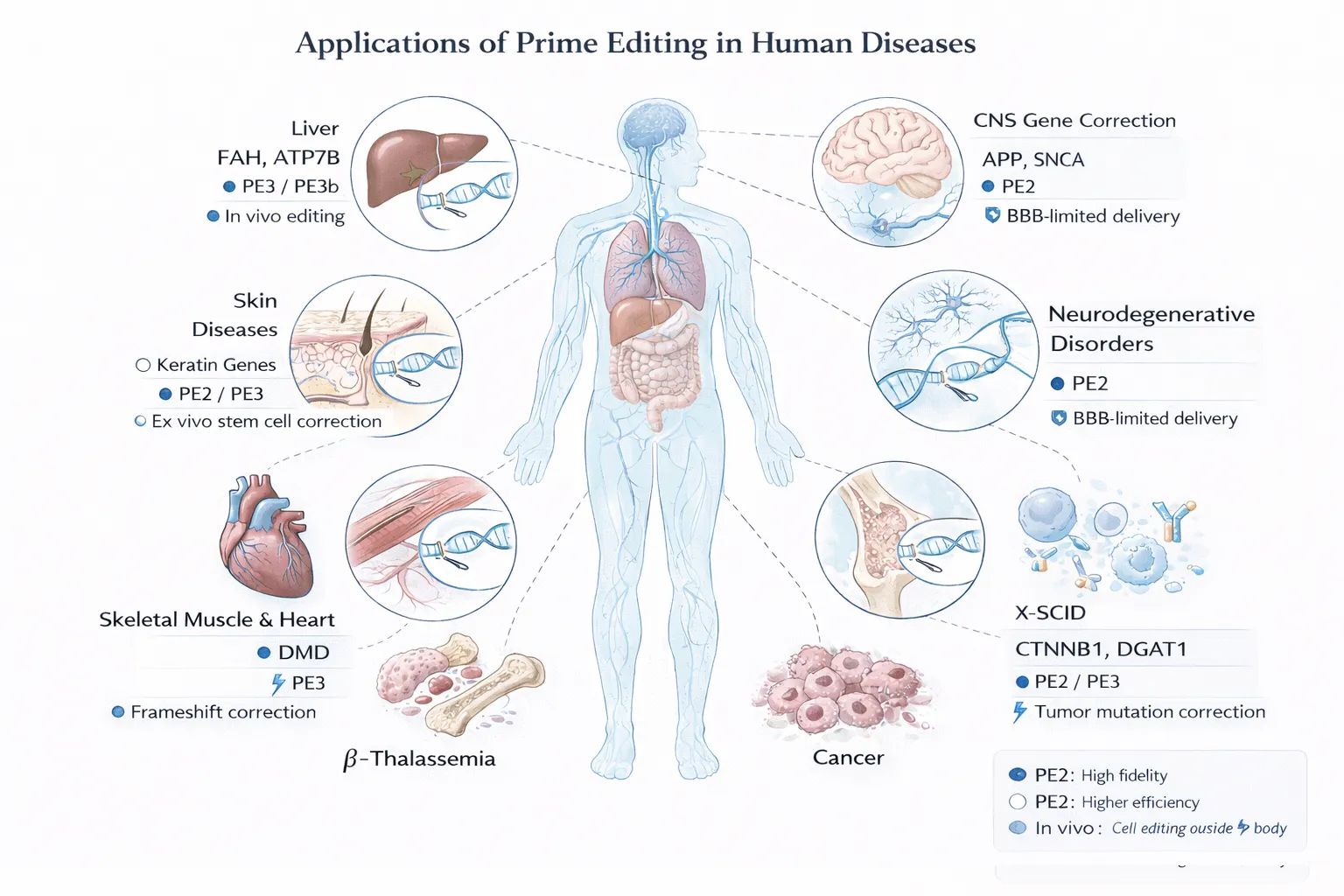
4. Prime Editing in Therapeutic Applications
Prime editing has rapidly progressed from a conceptual genome editing tool to a preclinical therapeutic platform, with demonstrated applications across multiple organ systems and disease classes. Its ability to precisely correct point mutations, small insertions, and deletions makes it particularly suitable for monogenic diseases, which constitute a large fraction of inherited disorders ([MDPI][1]).
Importantly, most current applications remain at the preclinical stage, including in vitro human cell systems and in vivo animal models, but they provide strong proof-of-concept for future clinical translation ([ScienceDirect][2]).
4.1 Application of Prime Editing to Liver Hereditary Diseases
Liver diseases represent one of the most advanced areas for prime editing applications due to:
- High accessibility via systemic delivery
- Strong regenerative capacity of hepatocytes
- Selective growth advantage of corrected cells
Case Study: Hereditary Tyrosinemia Type 1 (HT1)
Multiple studies have demonstrated successful correction of FAH gene mutations, which cause HT1:
- Prime editing corrected a G→A mutation in hepatic progenitor cells with ~2.3% efficiency
- After transplantation, corrected hepatocytes expanded to ~34% of the liver population, due to selective growth advantage ([MDPI][1])
In this study, in vivo editing achieved:
- Up to 11.5% genomic correction
- >60% FAH-positive hepatocytes, indicating functional recovery ([MDPI][1])
PEDAR Strategy (Prime Editing–Mediated Deletion and Repair)
- Enabled removal of large pathogenic insertions (~1.38 kb)
- Simultaneously repaired the locus with precise sequence replacement
- Demonstrated restoration of protein function in vivo ([MDPI][1])
Even modest editing efficiencies can yield therapeutic benefit due to clonal expansion of corrected hepatocytes, making the liver an ideal target for PE-based therapies.
4.2 Application of Prime Editing to Skin Disease
Skin diseases are particularly well-suited for ex vivo gene editing approaches, where cells can be edited outside the body and reimplanted.
Advantages:
- Direct accessibility of tissue
- Ability to expand corrected keratinocytes in vitro
- Reduced systemic delivery challenges
Applications:
- Correction of mutations in epidermolysis bullosa and keratin disorders
- Potential for long-term therapeutic benefit via stem cell-based skin regeneration
Although specific efficiency data are still limited, the MDPI review highlights skin as a promising early clinical target due to ease of manipulation and monitoring ([MDPI][1]).
4.3 Application to Skeletal and Cardiac Muscle Diseases
Prime editing has been explored in diseases such as:
- Duchenne muscular dystrophy (DMD)
- Inherited cardiomyopathies
Mechanistic Application
- Correction of frame-shift mutations or exon deletions
- Restoration of functional protein expression (e.g., dystrophin)
Experimental Evidence
- PE3-mediated editing successfully restored dystrophin expression in human iPSC-derived cardiomyocytes ([ScienceDirect][2])
Challenges
- Efficient delivery to large and distributed muscle tissues
- Achieving sufficient editing across multinucleated muscle fibers
Muscle diseases require system-wide delivery and high editing coverage, making delivery the primary bottleneck rather than editing capability.
4.4 Application to Neurodegenerative Diseases
Prime editing offers potential for correcting mutations underlying:
- Alzheimer’s disease
- Parkinson’s disease
- Other neurogenetic disorders
Technical Considerations
- Neurons are largely non-dividing, favoring PE over HDR-based editing
- Precision is critical due to sensitivity of neural tissue
Major Challenge: Delivery
One of the known challenges to not just Prime Editing and many other therapeutic approaches the Blood–brain barrier (BBB), which limits systemic access. So alternative solutions might be needed:
- AAV vectors with CNS tropism
- Nanoparticle-based delivery systems
Although still in early stages, PE is considered promising due to its low toxicity and high precision, which are essential for neurological applications ([MDPI][1]).
4.5 Application to Beta-Thalassemia
Beta-thalassemia is one of the most well-characterized targets for prime editing.
Disease Background
- Caused by mutations in the HBB gene
- Leads to defective β-globin production and severe anemia
Prime Editing Strategy
- Direct correction of disease-causing mutations
- Introduction of silent PAM-disrupting mutations to prevent re-editing
Key Findings
- ~14% correction efficiency in mouse models
- Elimination of disease phenotype (restoration of hematological function) ([MDPI][1])
- Some byproduct edits (~32%), highlighting need for optimization
- Alternative Strategy includes reactivation of fetal hemoglobin (HbF) as a compensatory mechanism
Prime editing enables precise correction of pathogenic alleles, offering a potentially curative approach compared to symptomatic treatments.
4.6 Application to X-Linked Severe Combined Immunodeficiency (X-SCID)
X-SCID is caused by mutations in the IL2RG gene, leading to severe immune dysfunction.
Prime Editing Applications
- Modeling disease mutations in vitro
- Correcting mutations in patient-derived T cells
Experimental Outcomes have demonstrated Editing efficiencies in the order of:
- ~31% in K562 cells
- ~26% in primary human T cells ([MDPI][1])
Challenges
- Limited expansion of corrected patient cells
- Presence of somatic mosaicism complicates therapeutic correction
Prime editing demonstrates strong potential for ex vivo immune cell therapies, though optimization of cell expansion and editing persistence is needed.
4.7 Application to Cancer
Prime editing is being explored in oncology through two major approaches:
1. Direct Tumor Genome Editing
- Correction of oncogenic mutations
- Targeting cancer-driving pathways
2. Immune Cell Engineering
- Enhancing CAR-T cell therapies
- Modifying immune checkpoints
Some of the advantages of Prime Editing over Traditional CRISPR are the Reduced risk of:
- Chromosomal rearrangements
- Large deletions
- Off-target mutagenesis
Prime editing’s precision makes it particularly valuable for engineering therapeutic cells, where genomic integrity is critical.
4.8 Summary of Approaches
Across all therapeutic domains, prime editing demonstrates:
- Precise correction of disease-causing mutations
- Reduced genomic damage compared to DSB-based editing
- Broad applicability across tissues
Key Limitation
- Delivery remains the central challenge across all applications
Translational Outlook
While still largely preclinical, the accumulated evidence suggests that prime editing is a highly promising platform for next-generation gene therapy, particularly for monogenic and rare diseases.
5. Discussion & Future Directions
Prime editing is transitioning from proof-of-concept to clinically relevant applications, but several hurdles remain.
Key Challenges
- Variable efficiency across cell types
- Delivery limitations
- Off-target and byproduct formation
Future Opportunities
- Integration with AI-driven pegRNA design
- Development of next-generation PE variants
- Improved in vivo delivery systems
- Expansion to large genomic insertions and complex edits
The literature suggests that PE could become a central platform for precision genome engineering, complementing or even replacing earlier CRISPR modalities in specific applications.
6. Conclusion
Prime editing represents a paradigm shift in genome editing, enabling precise and versatile DNA modifications with reduced risk compared to traditional CRISPR systems. Its ability to correct a wide range of genetic mutations positions it as a powerful tool for next-generation gene therapies.
While challenges remain—particularly in delivery and efficiency—ongoing technological advancements are rapidly accelerating its path toward clinical translation and therapeutic impact.
- Explore our in-depth guide to CRISPR and next-generation genome editing technologies
- Learn how multi-omics and gene editing are shaping the future of precision medicine