Introduction
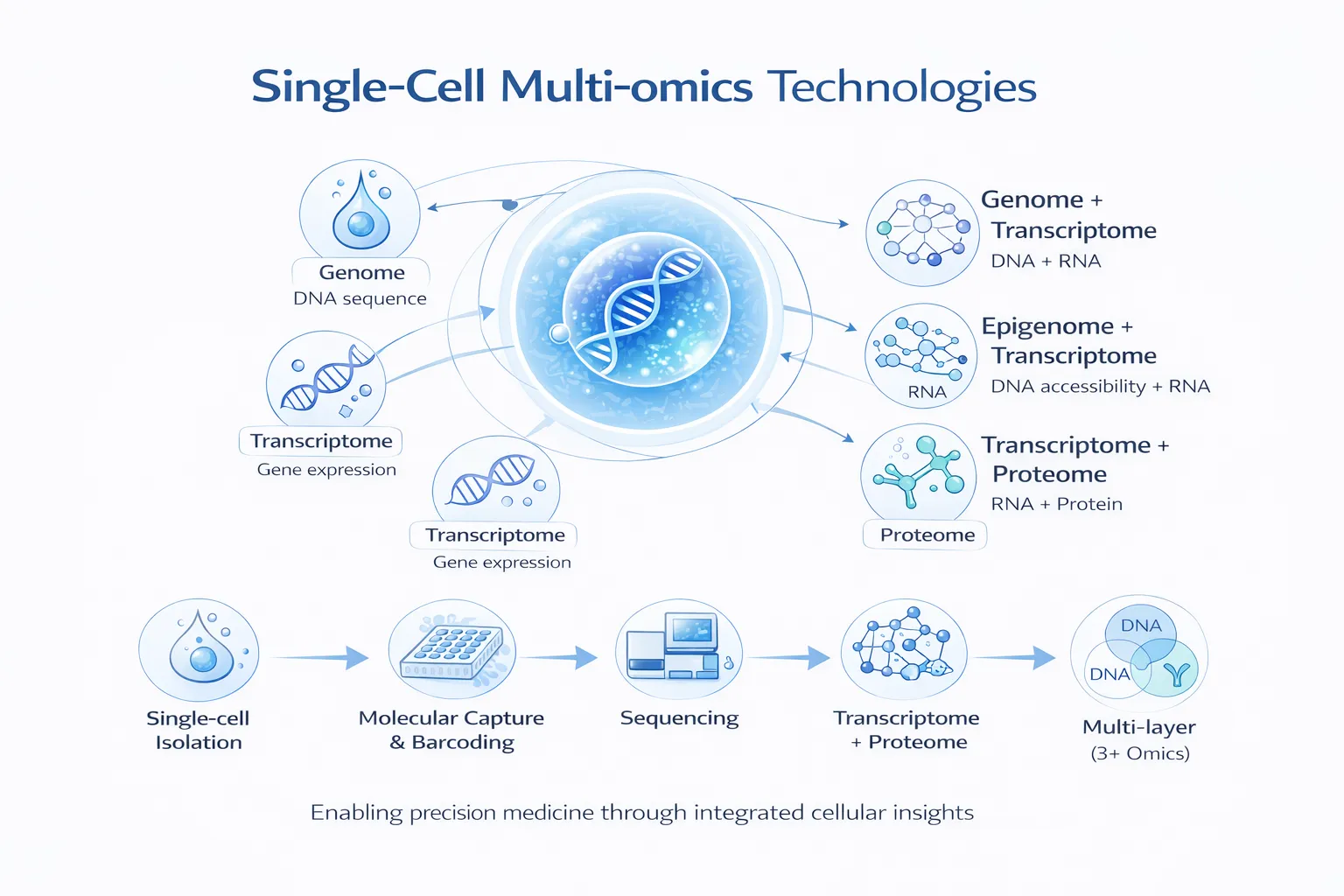
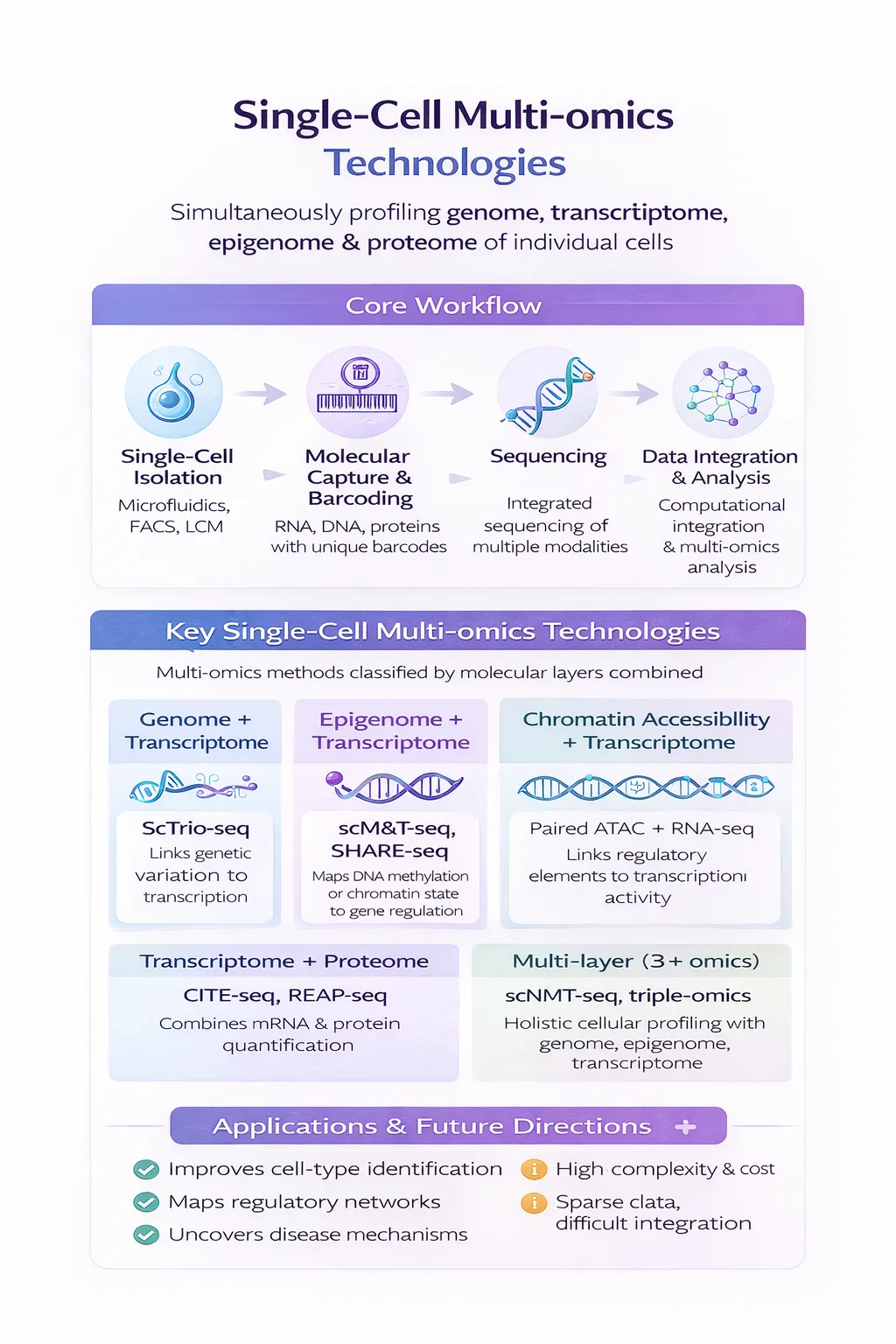
Single-cell multi-omics technologies are transforming how we understand biology—enabling researchers to simultaneously measure multiple molecular layers (genome, transcriptome, epigenome, proteome) within individual cells. This integrated view provides unprecedented insight into cellular heterogeneity, disease mechanisms, and regulatory biology.
Unlike traditional bulk or single-omics approaches, these methods capture interconnected molecular signals from the same cell, enabling direct inference of regulatory relationships and causal mechanisms across biological layers (Chappell et al., Annual Review of Genomics and Human Genetics).
What is Single-Cell Multi-omics?
Single-cell multi-omics refers to experimental and computational approaches that jointly profile multiple omics modalities—such as DNA mutations, RNA expression, chromatin accessibility, DNA methylation, and protein abundance—from a single cell.
These approaches allow researchers to:
- Link genotype to phenotype at cellular resolution
- Identify rare or transient cell states
- Infer gene regulatory networks
- Characterize disease heterogeneity
Recent advances emphasize integrating transcriptomic, epigenomic, and proteomic layers to better understand cellular identity and function (Lee et al., Experimental & Molecular Medicine, 2024).
🔬 Core Workflow of Single-Cell Multi-omics
Single-cell multi-omics experiments involve a multi-stage pipeline combining experimental innovation and computational integration.
1. Single-Cell Isolation
The first step is isolating individual cells while preserving molecular integrity. Here are some of the key technologies useful for single cell isolation:
Key technologies:
- Microfluidics (droplet-based systems): High-throughput encapsulation of cells into droplets (e.g., 10x Genomics platforms), enabling thousands of cells to be processed simultaneously.
- Fluorescence-Activated Cell Sorting (FACS): Enables selection of specific cell populations using surface markers, improving signal specificity.
- Microwell-based systems: Captures individual cells in wells for controlled processing.
- Laser capture microdissection (LCM): Preserves spatial context but operates at lower throughput.
👉 Insight: Trade-offs exist between throughput, sensitivity, and preservation of spatial/temporal context, which directly impact downstream multi-omics integration.
2. Molecular Capture & Barcoding
Once isolated, biomolecules are captured and labeled using various mechanisms. The following is a listing of the popular mechanisms:
Key mechanisms:
- Cell-specific barcoding: Assigns unique identifiers to molecules from each cell.
- Modality-specific tagging: Different molecular layers (RNA, DNA, proteins) are tagged with distinct barcodes or adapters.
- Antibody-based tagging (e.g., CITE-seq): Uses oligonucleotide-labeled antibodies to quantify proteins alongside RNA.
👉 Key challenge: Efficient co-capture of multiple molecular layers from the same cell without introducing bias or loss.
3. Library Preparation & Sequencing
Captured molecules are then converted into sequencing libraries, for further preparation and sequencing. Modern protocols integrate multiple library types into a single sequencing workflow, improving efficiency but increasing complexity. Some of the sequencing workflow is below:
- RNA → cDNA libraries
- Chromatin accessibility → ATAC-seq libraries
- DNA methylation → bisulfite sequencing
- Proteins → oligo-tag sequencing
4. Data Processing & Integration
Multi-omics datasets are inherently heterogeneous, sparse, and high-dimensional. So the real challenge is in the computational aspects of data normalisation and integration. The following is are some of the computational tasks in the data processing workflow:
- Data normalization across modalities
- Dimensionality reduction
- Batch correction
- Cross-modality alignment
Integration strategies:
Various experts believe that computational integration is now the primary bottleneck, not data generation. This is due to multi-model, heterogenous and sparse data sets in multi-omics. However, the following are broad strategies that can be used.
- Early integration (feature-level fusion)
- Late integration (model-level fusion)
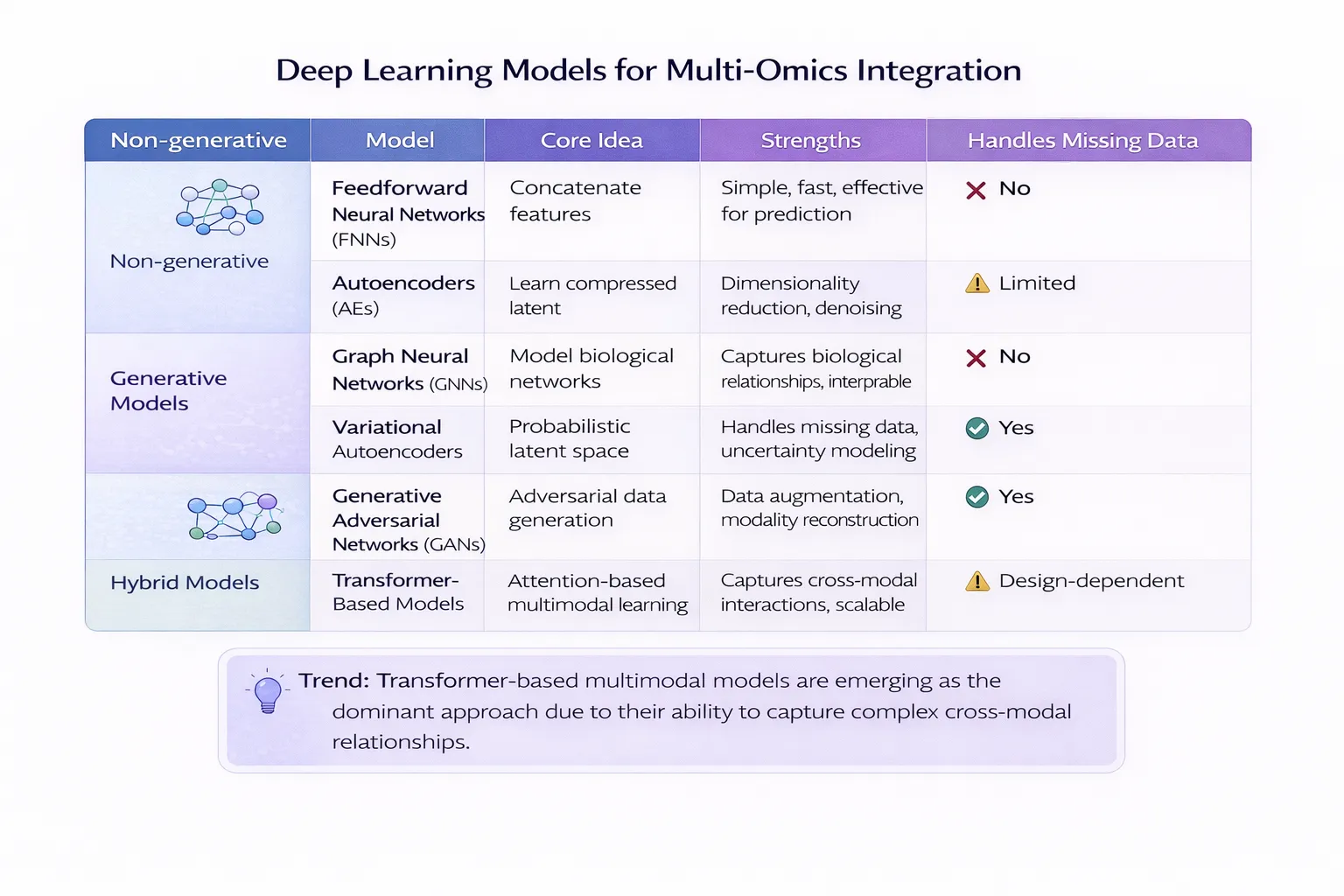
- Joint latent space learning (deep learning approaches)
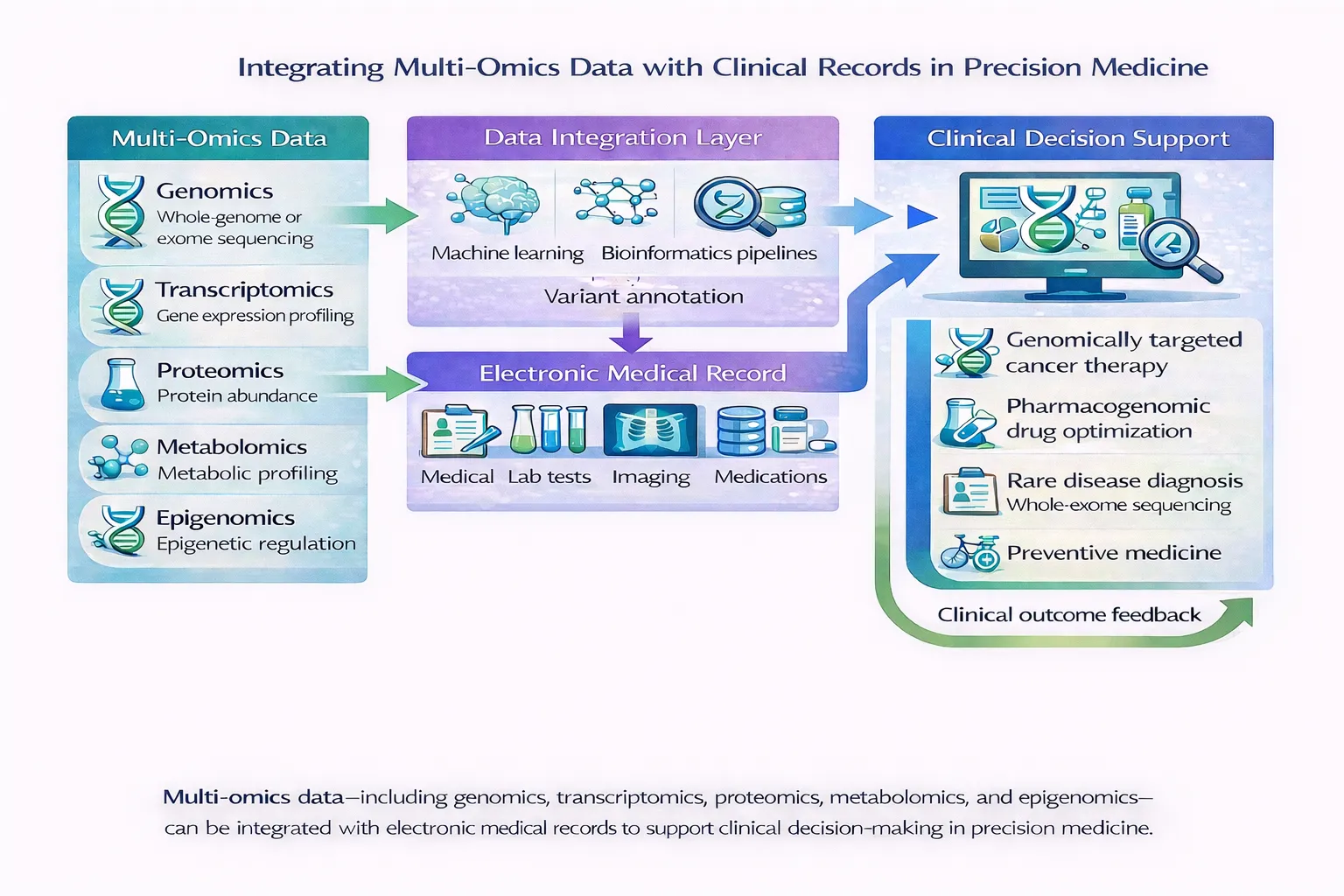
👉 You can learn more about how AI enhances multi-omics integration Read: Deep Learning for Multi-Omics Integration in Precision Medicine
5. Downstream Analysis
Once integrated, data enables:
- Cell-type identification
- Trajectory inference (developmental lineages)
- Gene regulatory network reconstruction
- Multi-layer biomarker discovery
🧬 Major Categories of Single-Cell Multi-omics Technologies
Single-cell multi-omics technologies are classified based on which molecular layers are jointly profiled.
1. Genome + Transcriptome Technologies
These technologies simultaneously measure DNA mutations (genome) and RNA expression (transcriptome) from the same cell.
Examples of these technologies include scTrio-seq and G&T-seq (Genome & Transcriptome sequencing)
Their key contribution is they directly link genetic variation to transcriptional output.
Use cases:
- Cancer heterogeneity
- Clonal evolution
- Mutation impact analysis
👉 Limitation: DNA and RNA require different preparation workflows, making co-assay technically challenging.
2. Epigenome + Transcriptome Technologies
These approaches measure DNA methylation OR chromatin accessibility and RNA expression within the isolated cells. They are often valuable in revealing how epigenetic regulation controls gene expression. This is informative, because:
- Chromatin accessibility reflects regulatory potential
- DNA methylation reflects long-term gene silencing
Examples:
- scM&T-seq (methylation + transcriptome)
- SHARE-seq (chromatin accessibility + RNA)
- Paired scATAC-seq + scRNA-seq
👉 Challenge: Epigenomic data is highly sparse, requiring advanced computational modeling.
3. Transcriptome + Proteome Technologies
These technologies combine RNA sequencing and Protein quantification via antibody-derived tags. This allows scientists to bridge gaps between gene expression and functional protein levels. Their primary advantages are:
- Improved cell-type classification
- Captures post-transcriptional regulation
Examples:
- CITE-seq
- REAP-seq
👉 Limitation: Restricted to pre-selected protein panels.
4. Chromatin Accessibility + Transcriptome
These technologies simultaneously profile Open chromatin regions (ATAC-seq) and RNA expression. This enables the linkage of regulatory elements (enhancers/promoters) to gene expression, which are critical for
- Gene regulatory network reconstruction
- Developmental biology
Examples:
- SNARE-seq
- SHARE-seq
5. Multi-layer (Triple-omics and Beyond)
We use multi-layer technologies when we want to simultaneously capture the Genome, the Epigenome and the Transcriptome. This provides a holistic view of cellular regulation. However it comes with the trade off of:
- Lower throughput
- Higher cost
- Increased computational complexity
Examples:
- scNMT-seq (nucleosome positioning + methylation + transcription)
- Emerging triple-omics platforms
📊 Comparative Table of Technologies
| Technology Type | Molecular Layer Targeted | Modalities Measured | Key Idea | Strengths | Limitations | Applications |
|---|---|---|---|---|---|---|
| Genome + Transcriptome | Genetic variation | DNA + RNA | Link mutations to expression | Direct genotype–phenotype mapping | Complex workflows | Cancer evolution, lineage tracing |
| Epigenome + Transcriptome | Gene regulation | DNA methylation / chromatin + RNA | Study regulatory control | Reveals gene regulation | Sparse data | Developmental biology |
| Chromatin Accessibility + Transcriptome | Regulatory elements | ATAC + RNA | Link chromatin state to transcription | Captures regulatory dynamics | Computationally intensive | Gene regulation |
| Transcriptome + Proteome | Functional expression | RNA + proteins | Combine transcript and protein signals | Better cell typing | Limited protein panels | Immunology |
| Multi-layer (3+ omics) | Systems-level regulation | DNA + RNA + epigenome | Holistic cellular profiling | Most comprehensive | Costly, low throughput | Precision medicine |
🚀 Key Advantages of Single Cell Multi-omics
- Multi-layer biological insight
- Improved resolution of cellular heterogeneity
- Enhanced disease mechanism discovery
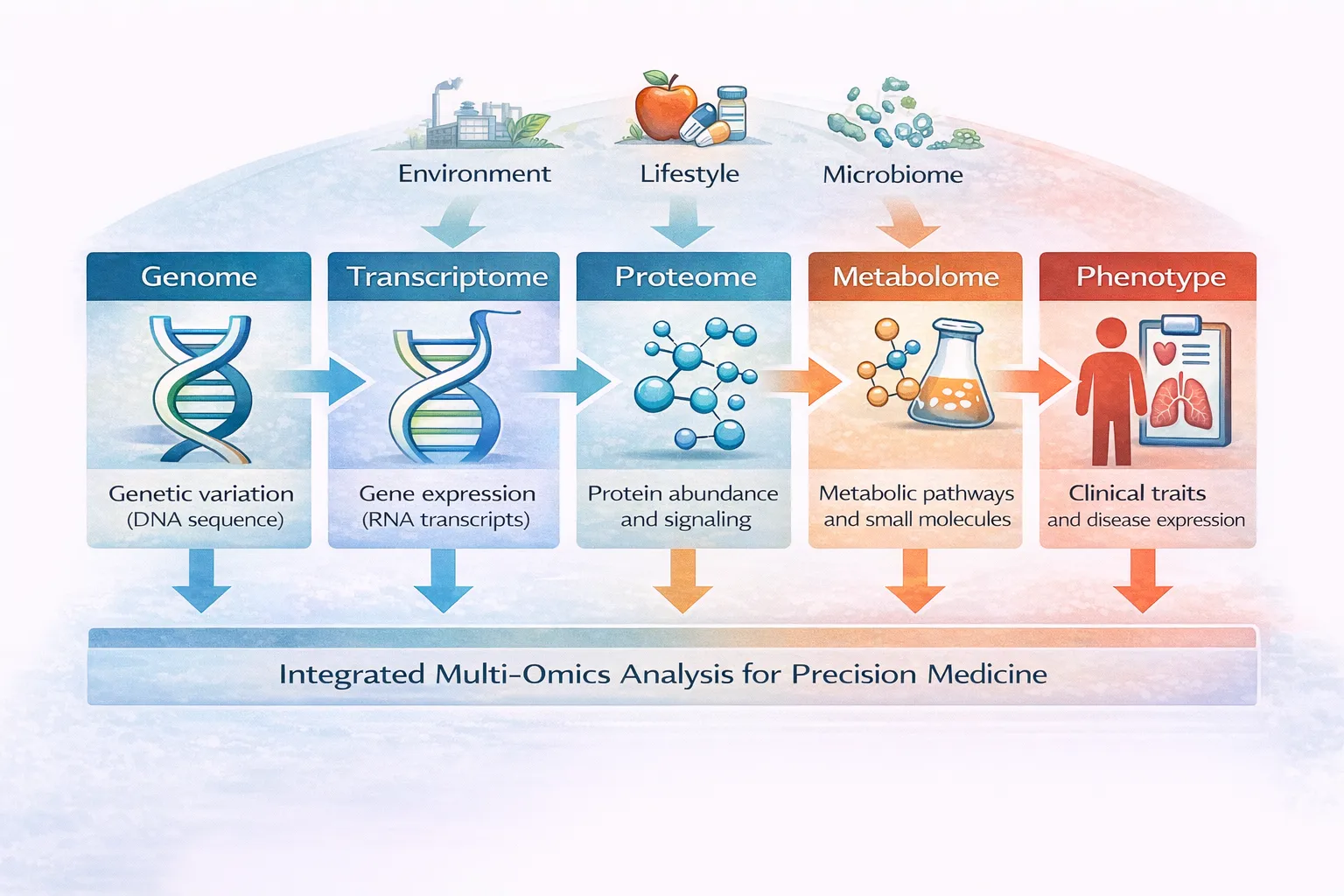
- Strong foundation for precision medicine
⚠️ Challenges
- Experimental complexity
- Data sparsity
- Integration difficulty
- Cost and scalability
🔮 Future Directions
While not covered by this article, there is a dimension of spatial context that can be considered relevant to single-cell multi-omics. This dimension recognises that within the human body, often single cells are part of a tissue or organ and have adjoining cells that might also have a role. So integration with spatial transcriptomics is an area of interest for multi-omics. Other future directions include
- AI-driven multimodal learning
- Clinical translation into diagnostics
- Standardization of protocols and pipelines
Conclusion
Single-cell multi-omics technologies are redefining biological research by enabling multi-layer, single-cell resolution insights. As experimental and computational methods continue to evolve, these technologies will play a central role in precision medicine, systems biology, and translational research.